goals and approach/
The Brain Science Management team of the Allen Institute's brain science accelerator leads our foundational scientific research in deciphering the cellular and circuit organization of the mammalian brains, from mouse to human, and providing open-access resources of data, knowledge and tools to accelerate neuroscience discoveries across the field. We do this by setting strategic goals, leading the development of new technology platforms and pipelines, leading the generation and public dissemination of comprehensive, multimodal datasets characterizing brain cell types and circuit networks, managing scientific operations, and engaging in collaborations with external researchers.
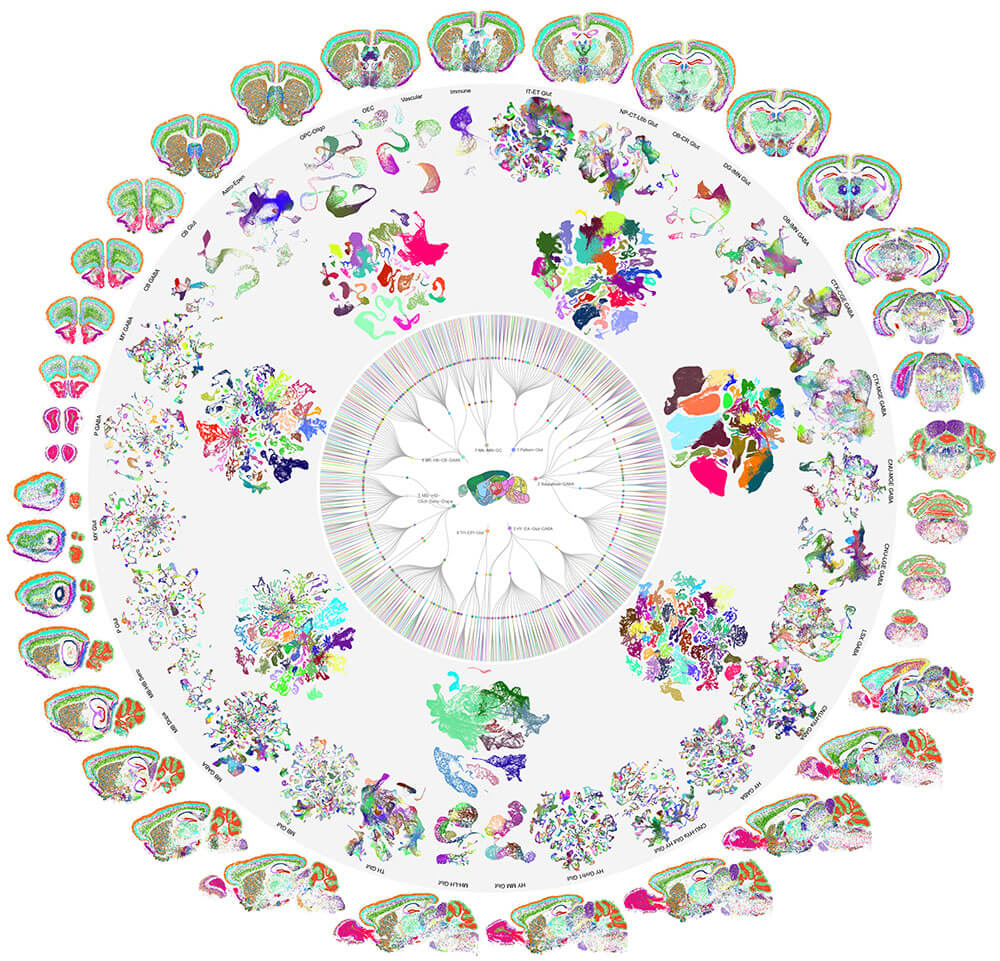
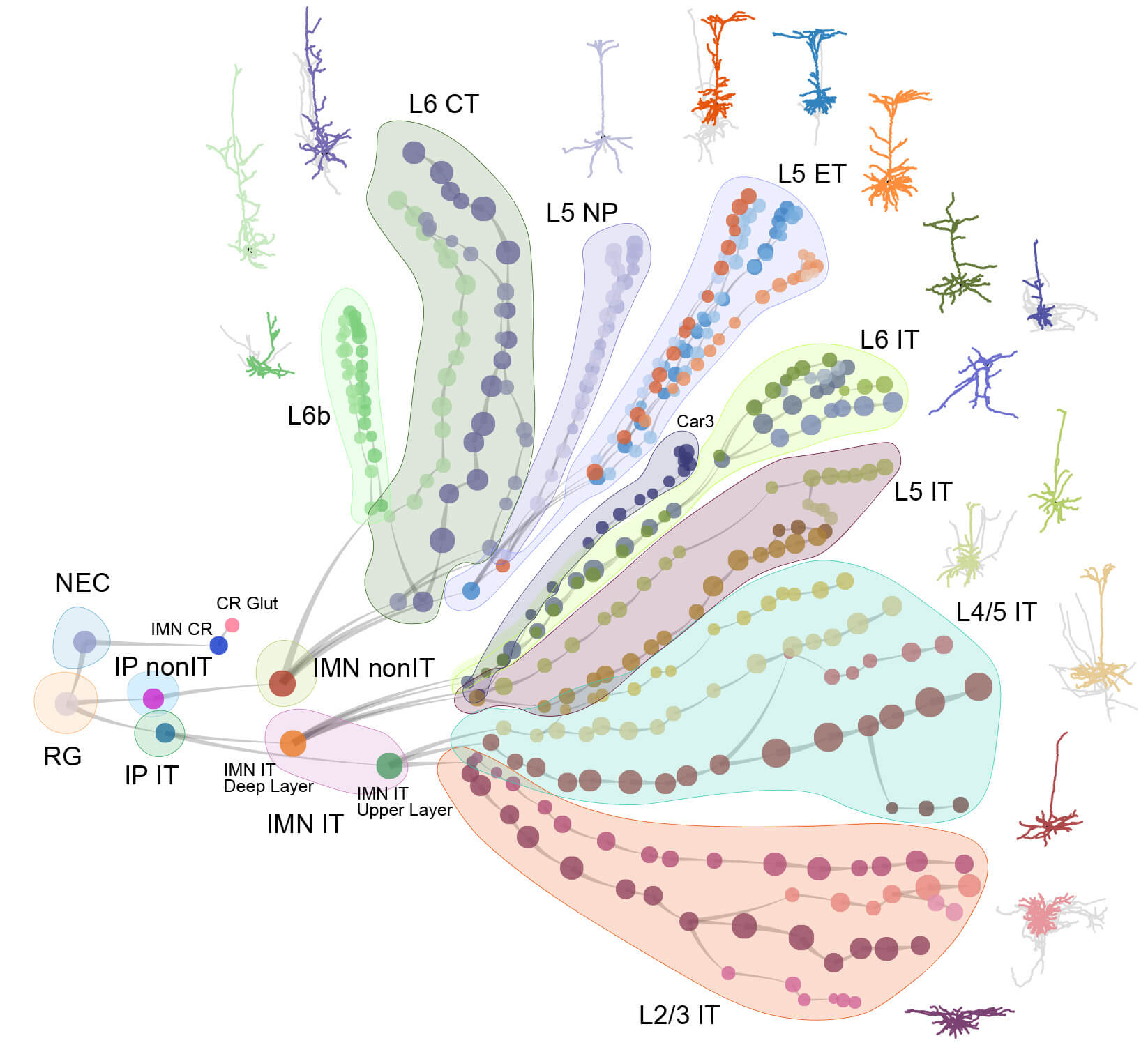
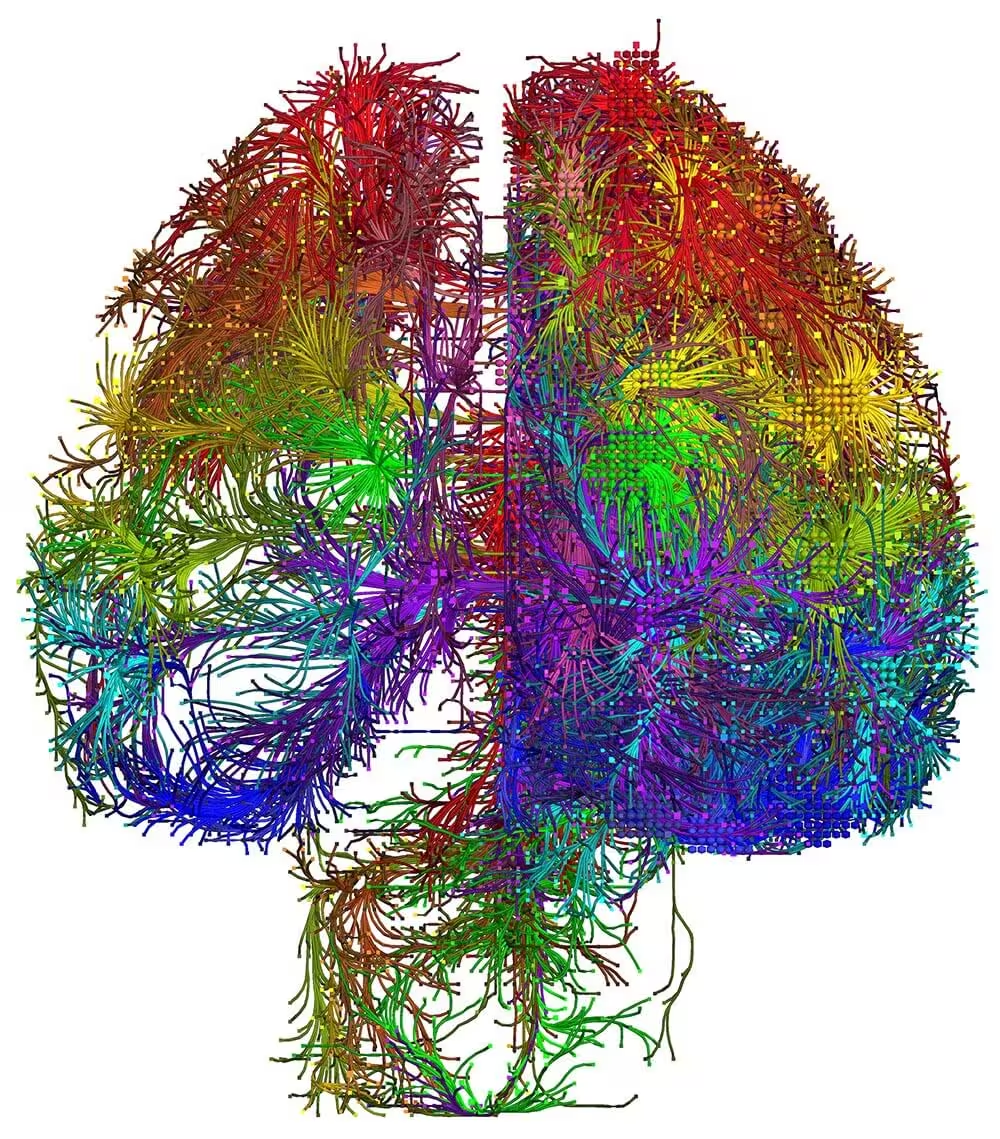
The mammalian brain is our most complex and mysterious organ, comprising millions to billions of cells and orchestrating our behaviors, emotions, cognition, and metabolism. To understand the function of the brain and how its dysfunction leads to brain diseases, it is essential to uncover the cell type composition of the brain, how the cell types are connected with each other and what their roles are in circuit function. At the brain science accelerator, we have built multiple technology platforms, including single-cell transcriptomics and multiomics, spatial transcriptomics, single and multi-patching electrophysiology, 3D reconstruction of neuronal morphology, brain-wide connectivity mapping, and synaptic-level connectomics by electron microscopy to characterize the molecular, anatomical, physiological, and connectional properties of brain cell types in a systematic manner, towards the creation of multi-modal cell atlases for the mouse, non-human primate and human brains. Our studies reveal extraordinary cellular diversity and underlying principles of brain organization. They establish foundational resources for deep and integrative investigations of cellular and circuit function, development, and evolution of the mammalian brain.
Scientists in the Brain Science Management department also carry out focused, cross-disciplinary research projects to gain a deeper understanding of the relationship between cell types’ molecular identities and connectional properties, and how cell types respond in various behavioral, pharmacological and diseased conditions.