goals and approach
The Human Cell Types team at the Allen Institute's brain science accelerator aims to characterize the diverse cell types of the mammalian brain and to build foundational open-access cell type classifications. This team applies cutting-edge single cell molecular genetic approaches to measure the gene expression signatures of hundreds of thousands or even millions of individual brain cells. They then apply computational approaches to identify cell type clusters and compare these cell types across individuals and species to understand how they change in health and disease and across evolutionary scale.
Additional focuses for this team are: defining the shapes, electrical properties and local connections of neuronal cell types in the same contexts, and creating viral genetic tools that allow observation and manipulation of specific brain cell populations to facilitate cell type characterization across species and to create new AAV therapeutics.

recent news
comparative brain cell atlasing projects
Alzheimer's disease projects
genetic tools and gene therapy projects
data and tools/
The Human Cell Types team has played driving roles in several public resources available on Allen Brain Map. These include:
Cell types in adult human brain
- Single nucleus RNA-sequencing (gene expression data)
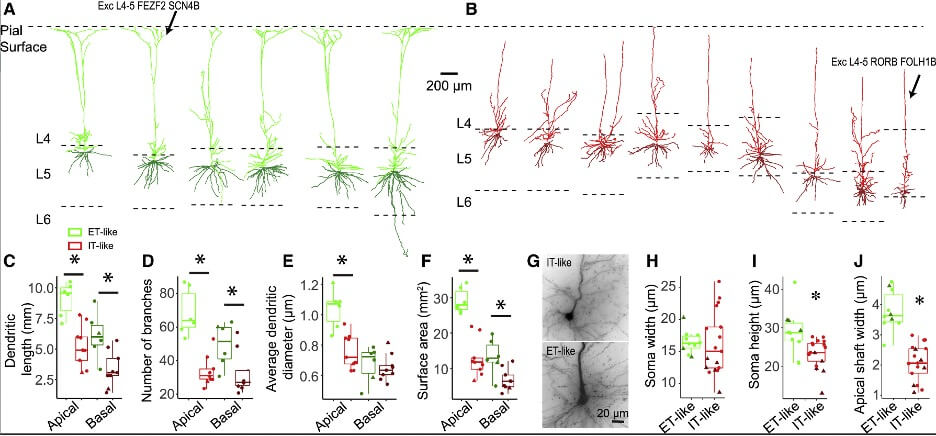
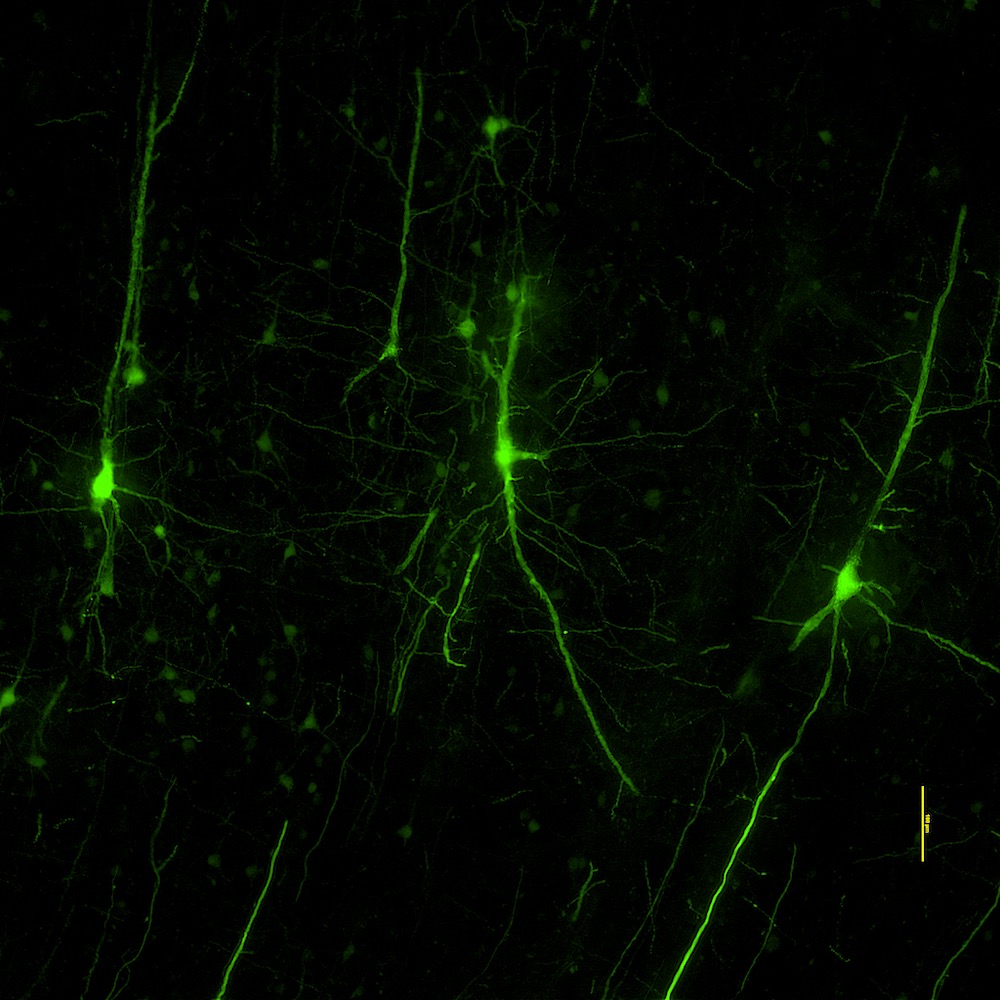
- Morphology and electrophysiology
- Patch-sequencing (multimodal characterization)
- Synaptic physiology (in vitro characterization)
- Cell Type Knowledge Explorer
- Explore: Cell Taxonomies
Gene expression atlases
- Adult human brain
- Developing human brain
- Adult and developing nonhuman primate
- Explore: Transcriptional Landscape of the Brain
Disease atlases
- Seattle Alzheimer’s Disease Brain Cell Atlas (SEA-AD)
- SEA-AD: Spatial transcriptomics
- SEA-AD: MapMyCells
- Aging, Dementia, and TBI Study
- IVY Glioblastoma Atlas Project
Genetic tools
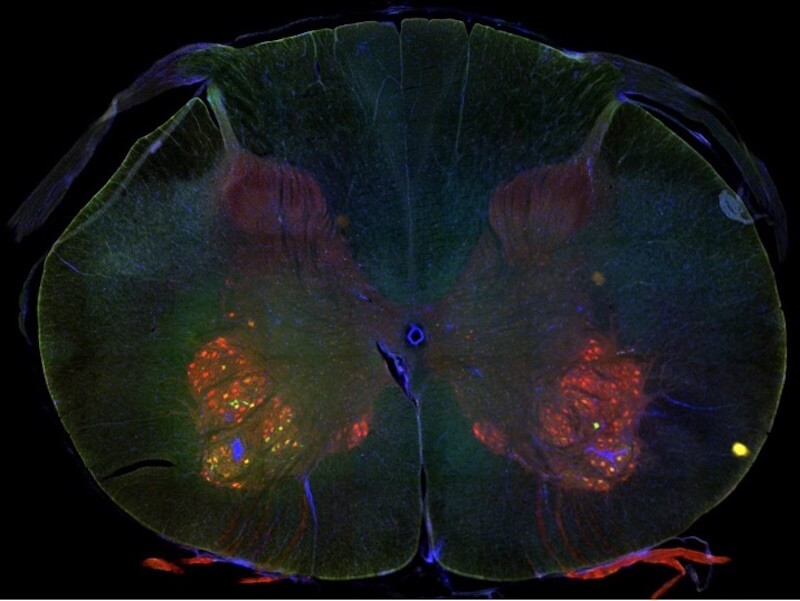
Neuroanatomy
Protocols