discover/
brain science
brain science
building a complete, interactive blueprint of the brain's cells and connections to unlock how it all works
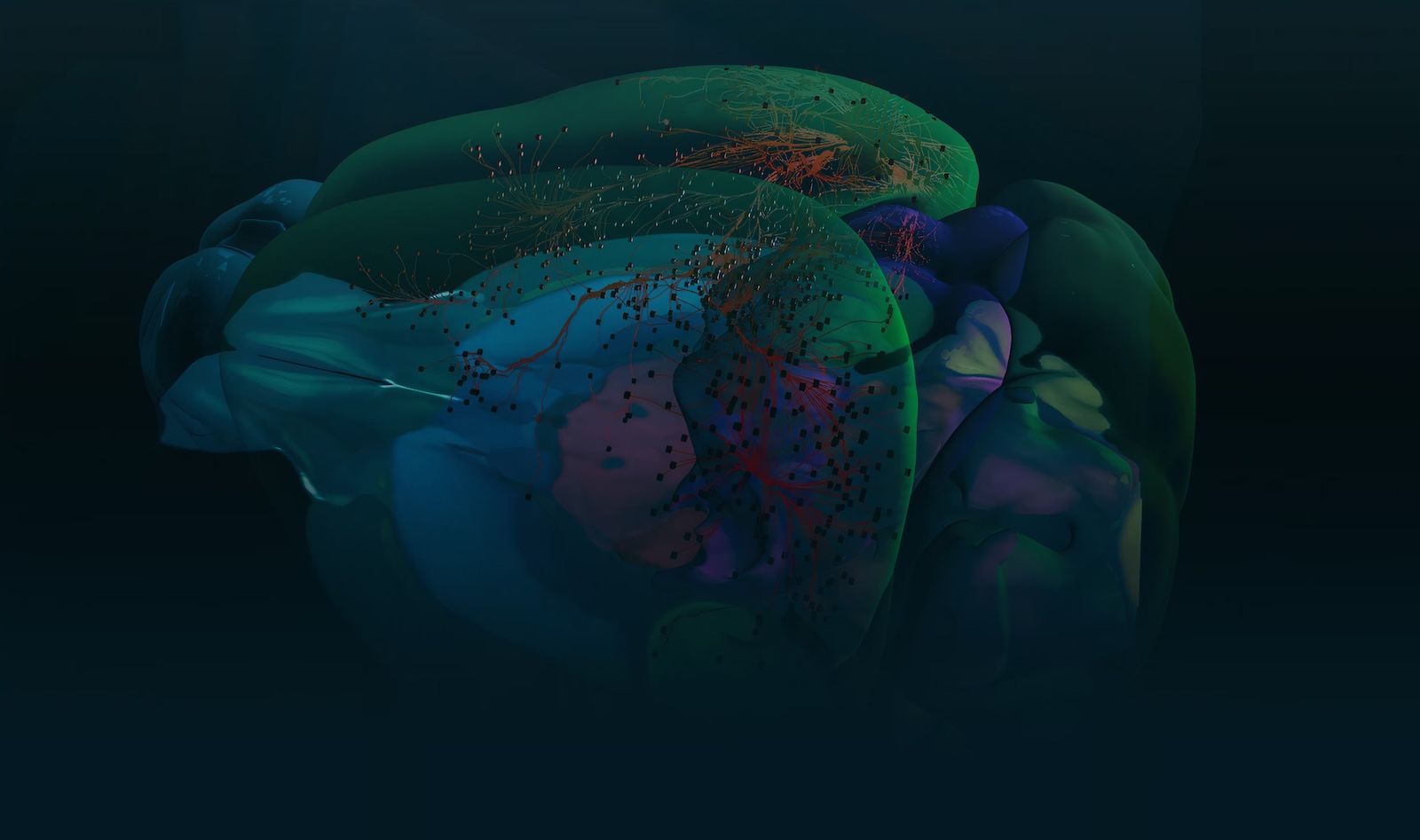

The brain's complexity is staggering. Understanding it requires a blueprint. We're building one.
Imagine trying to understand a city without a map: no street grid, no neighborhood names, no guide to which utilities connect where. That's the challenge brain scientists have faced for decades. We're creating the maps that change everything.
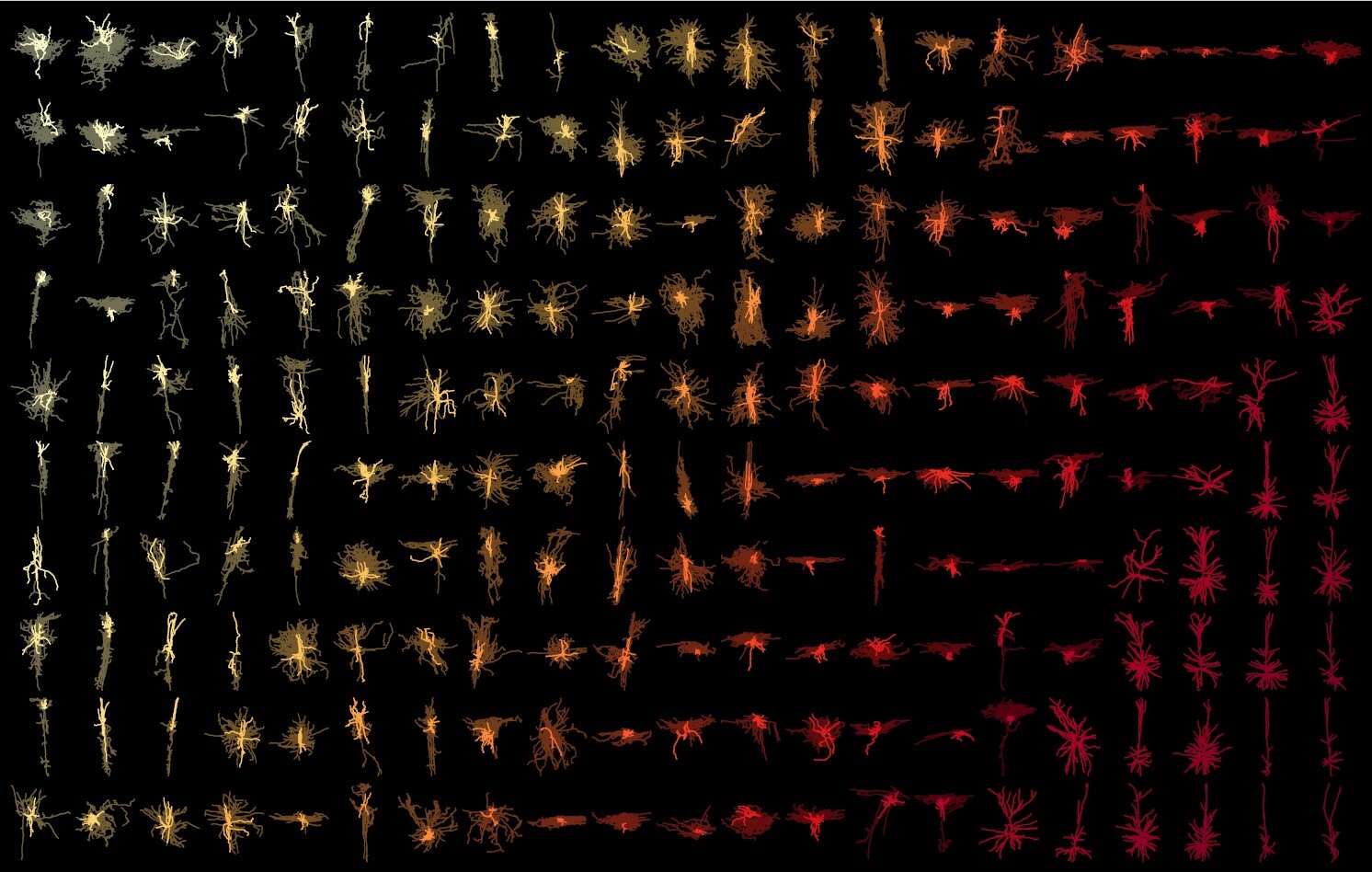
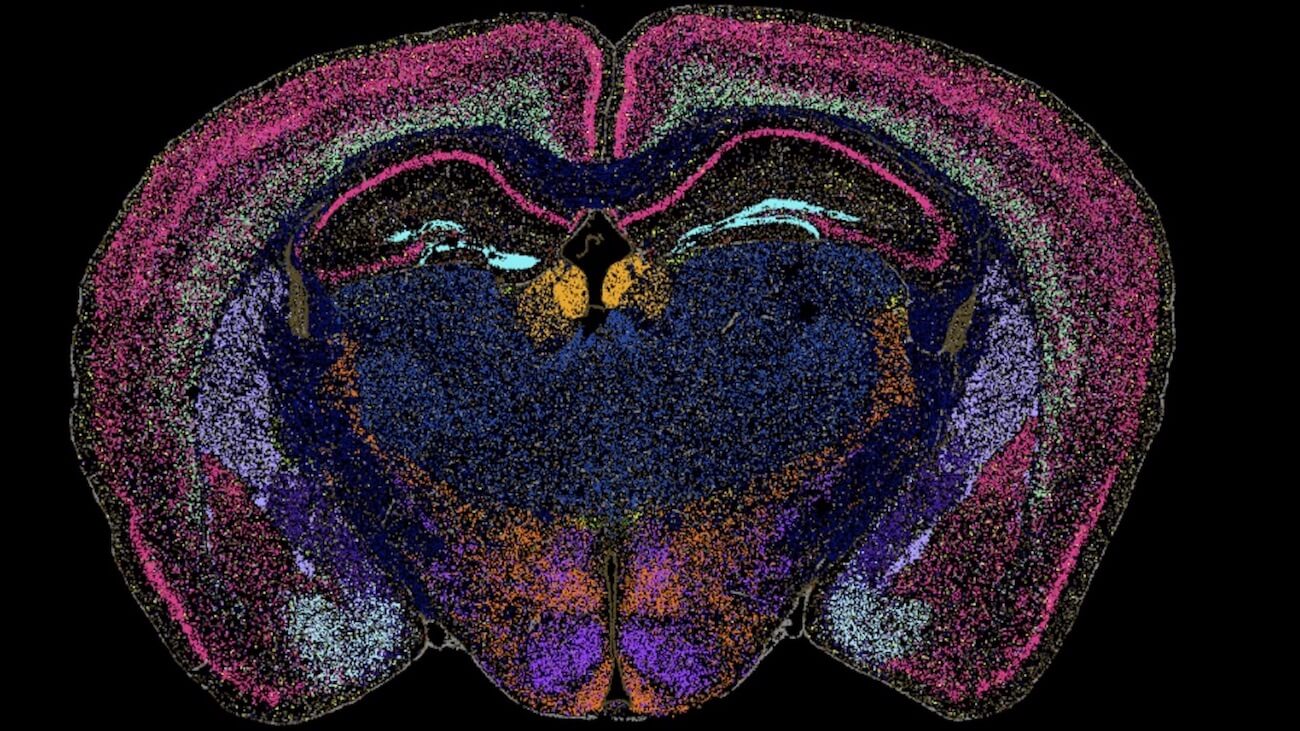
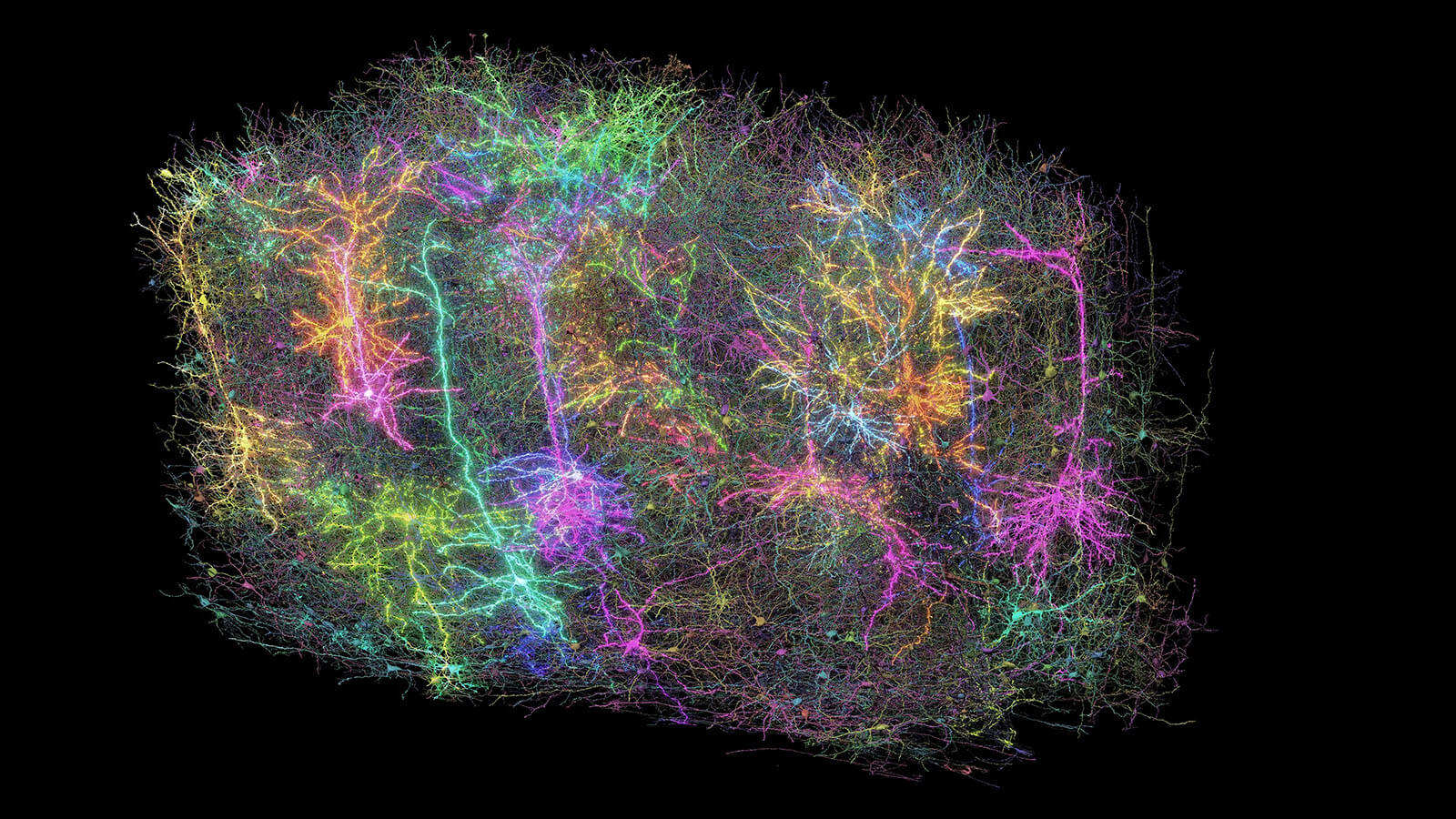
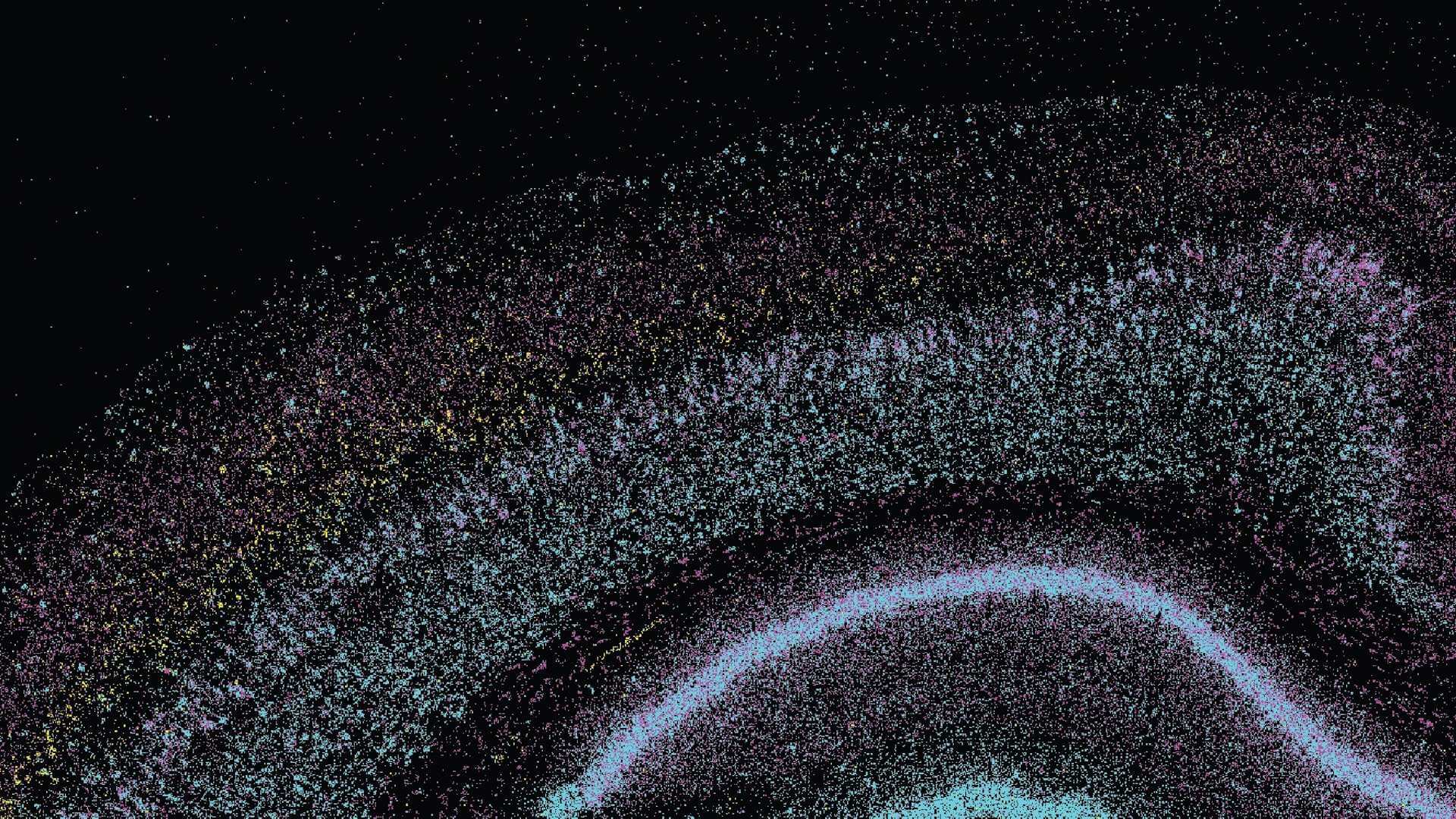
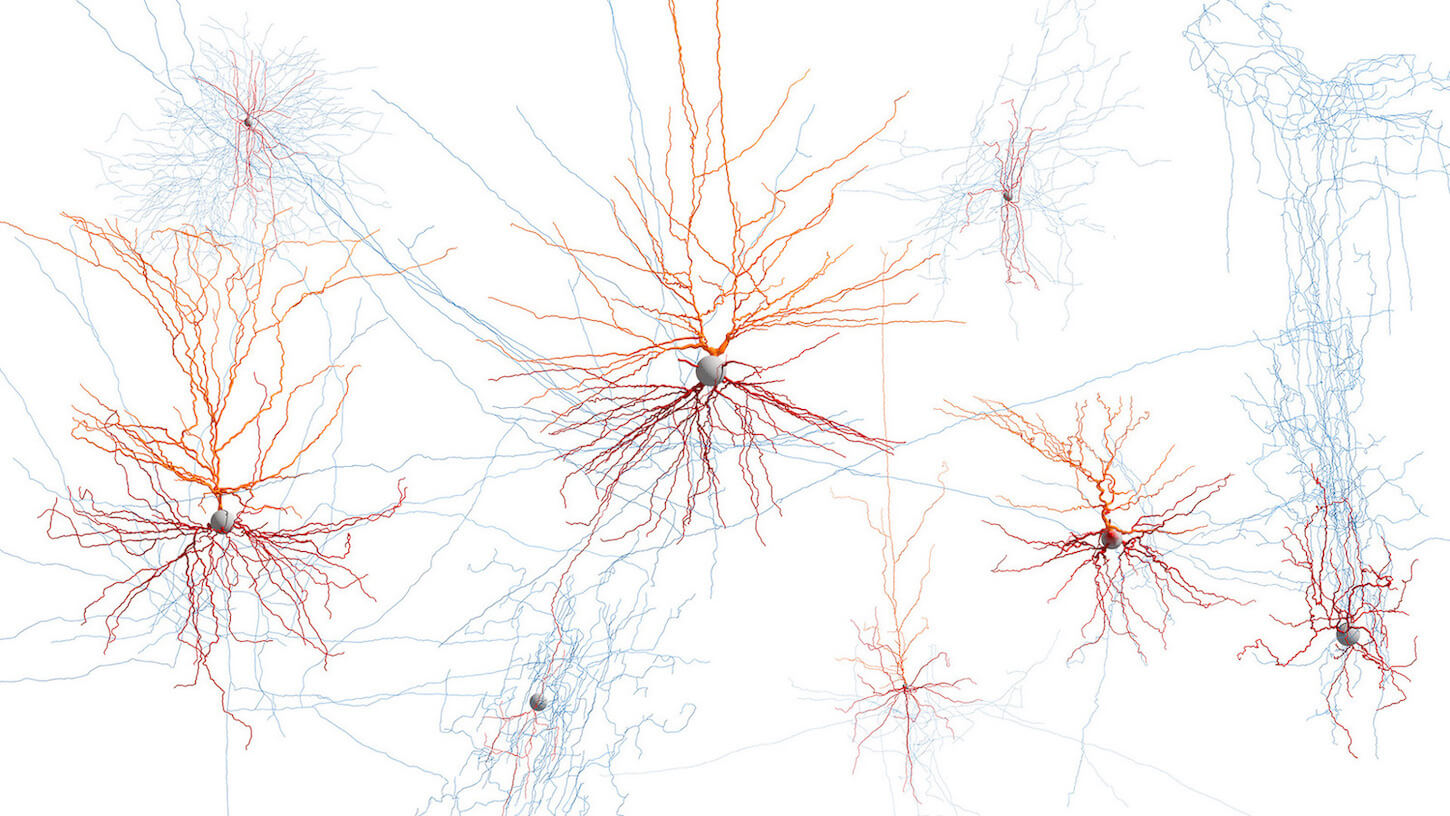
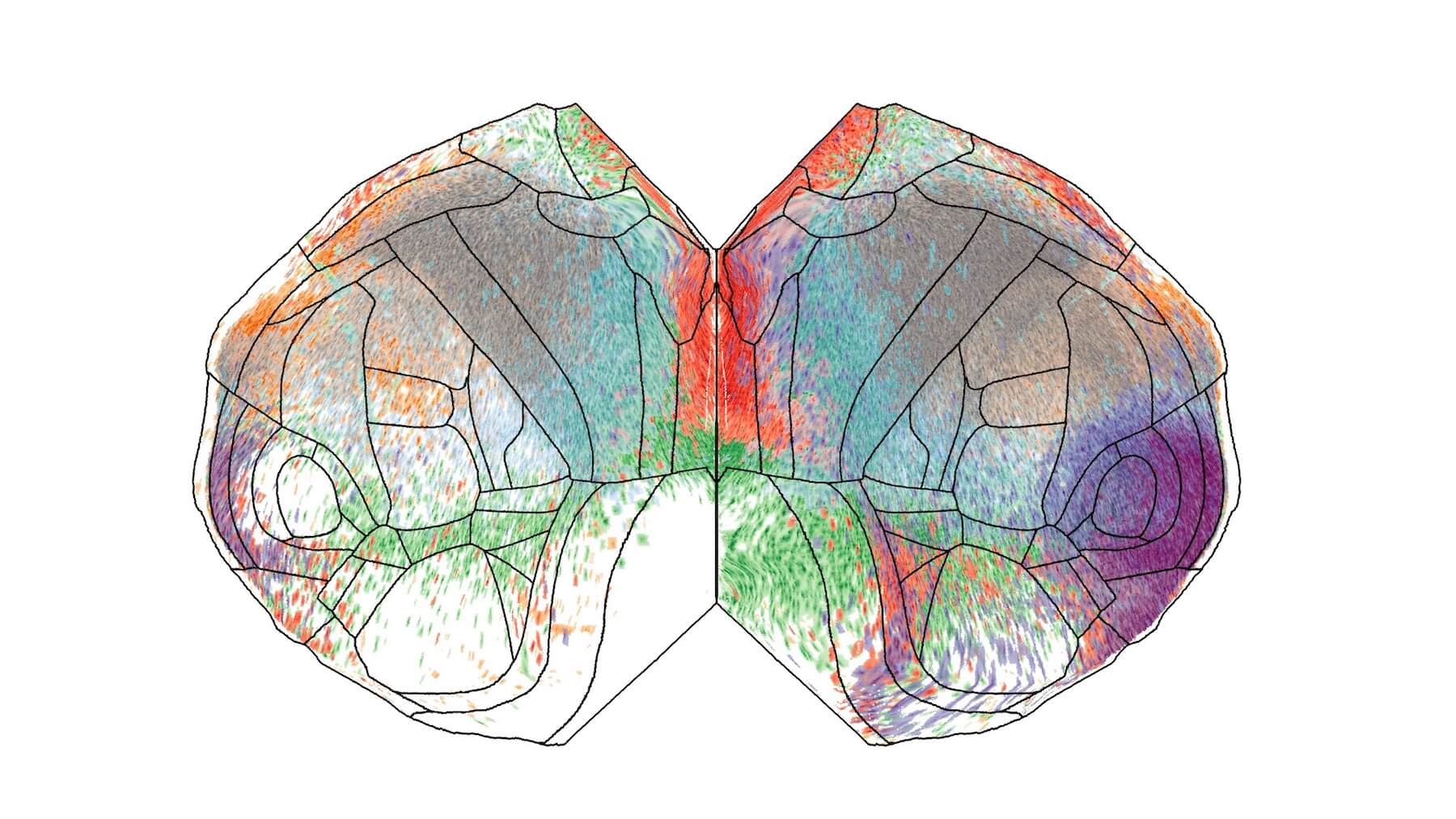
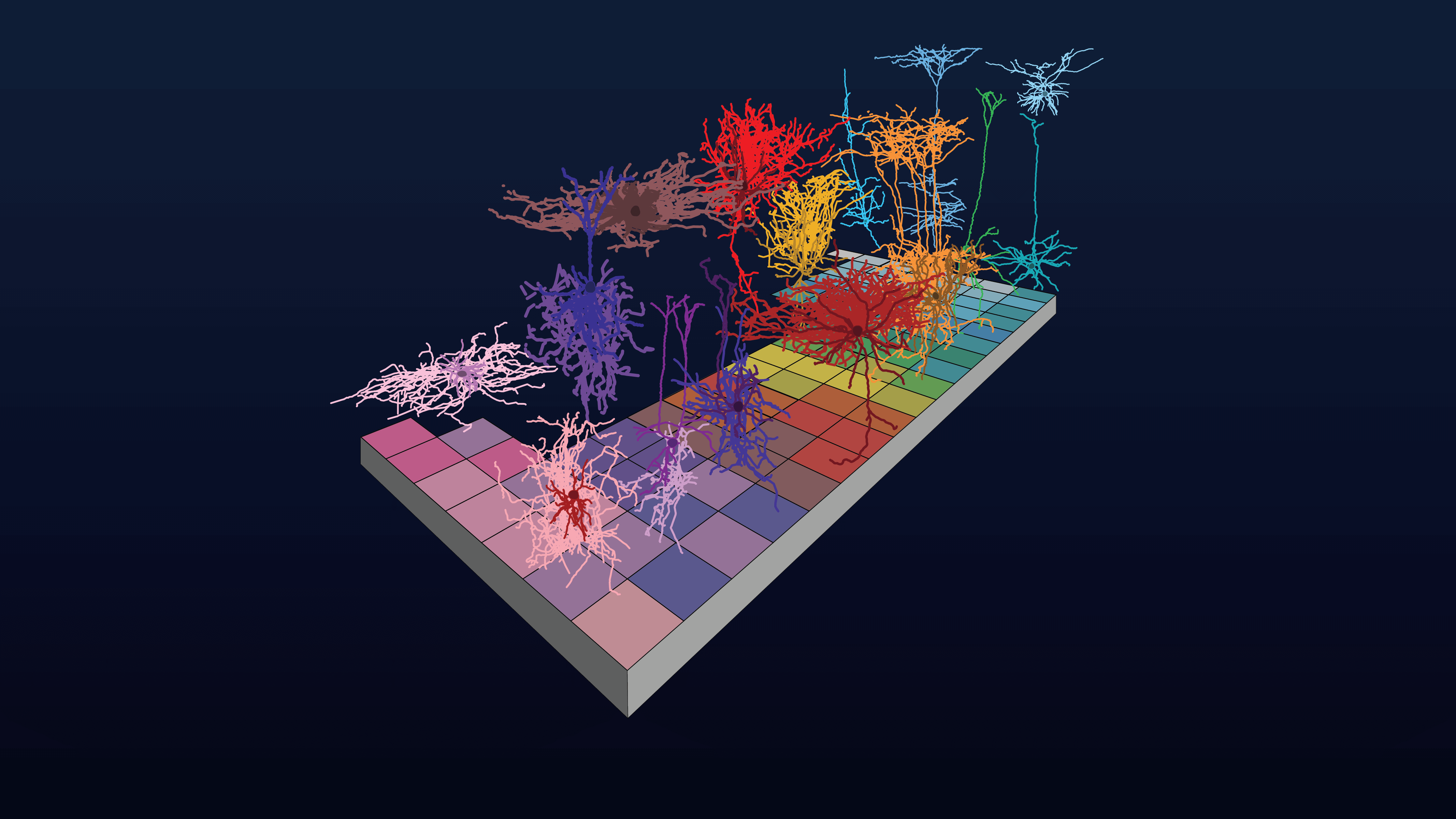
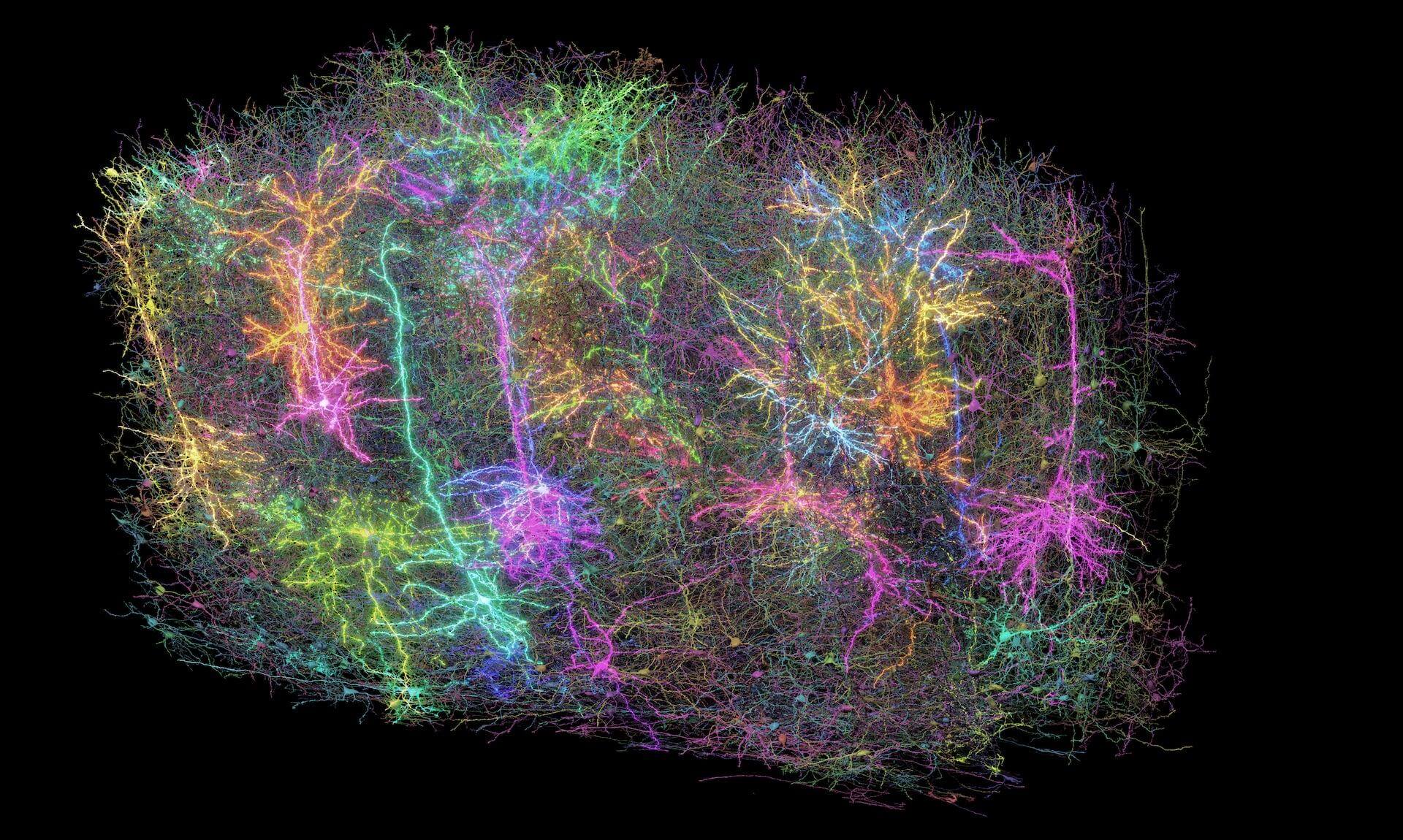
Our Brain Science team produces three interlocking resources: cell type atlases that catalog every kind of brain cell across ages and species, connectomes that chart the brain's intricate wiring, and a knowledge platform that knits it all together with powerful analysis tools. Every dataset is freely shared, so discoveries made here fuel discoveries everywhere.
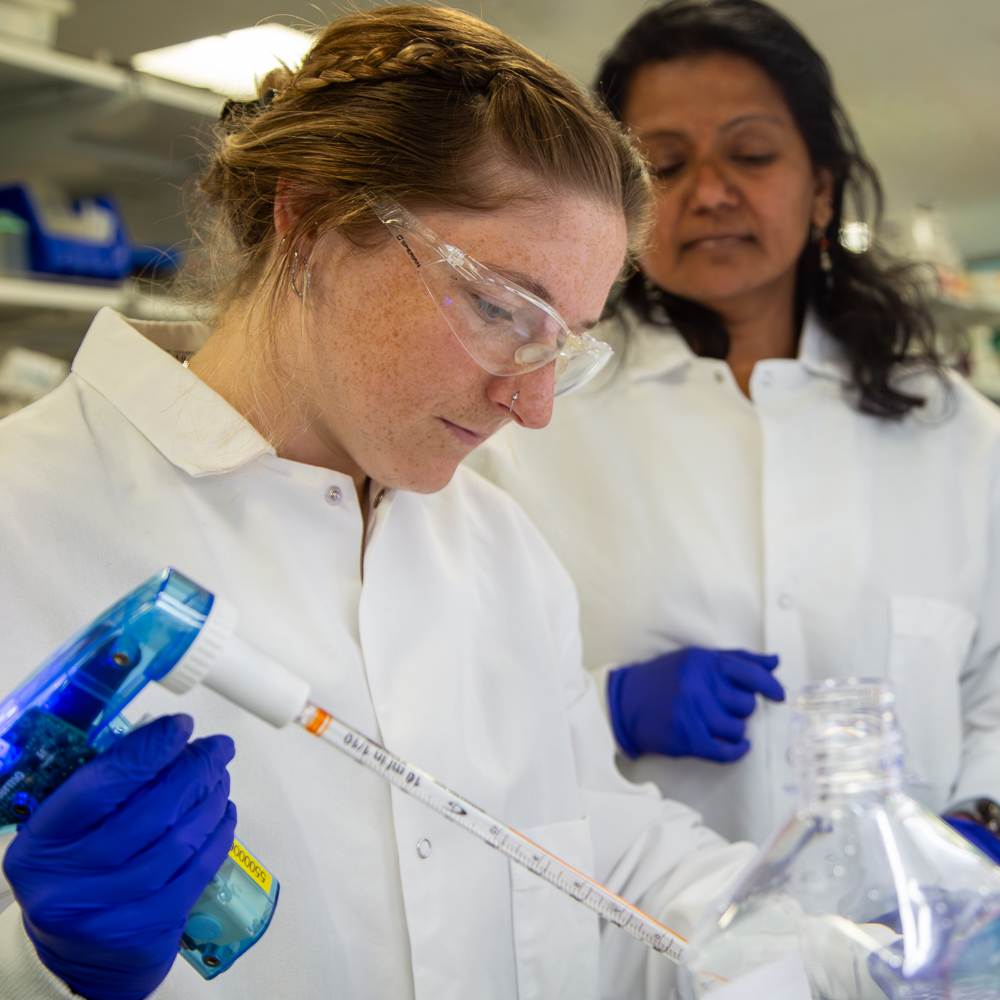

Our brain cell atlases, connectomes, and open knowledge platform give scientists worldwide the reference tools they need to accelerate every corner of brain research.
brain science news

By examining brain cell types and connections across species at a refined level and at the most comprehensive scale – we are working towards understanding the structure and function of the human brain itself.
Hongkui Zeng
Executive Vice President and Director, Brain Science

Block Quote
This is some text inside of a div block.
This is some text inside of a div block.
featured publications
access open tools, brain atlases, neuroscience databases, and more
brain science events

2026 Allen Institute Modeling Software Workshop
Join us at the 2026 Allen Institute Modeling Software Workshop for a hands-on opportunity to learn how to use open-source software modeling tools developed by the Allen Institute and our collaborators at the University of Illinois Urbana-Champaign (UIUC).
brain science advisory council