in this article
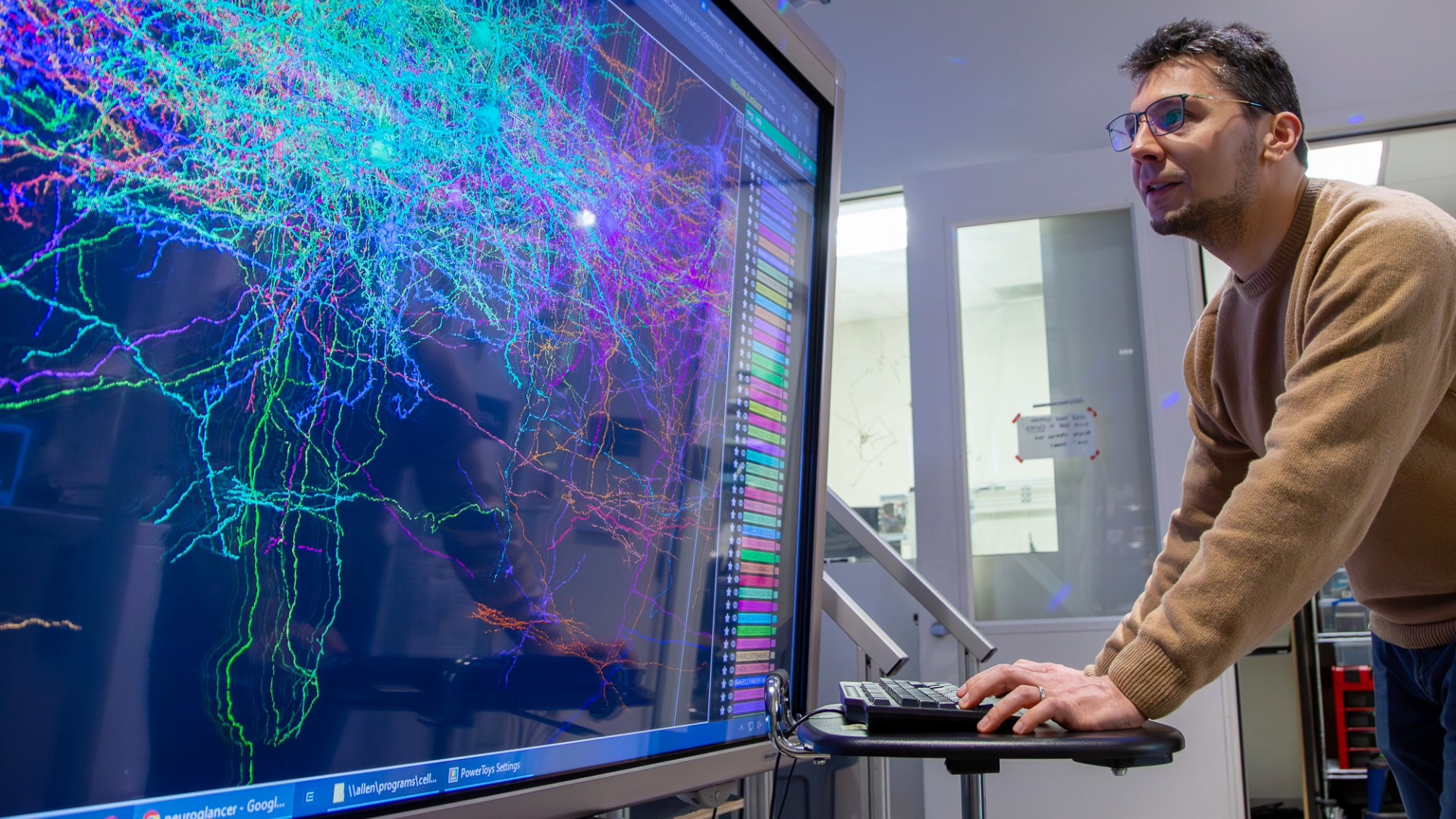
Scientists at the Allen Institute are launching the brain equivalent of the Human Genome Project, leading a new global collaboration to map the approximately 200 billion cells in the human brain by their type and function.
The collaboration is funded by the National Institutes of Health’s Brain Research Through Advancing Innovative Neurotechnologies® (BRAIN) Initiative as part of The BRAIN Initiative® Cell Atlas Network, or BICAN, and will also build detailed atlases of macaque and marmoset brains. Led by Ed Lein, Ph.D., Senior Investigator at the Allen Institute for Brain Science, a division of the Allen Institute, and Hongkui Zeng, Ph.D., Executive Vice President and Director of the Allen Institute for Brain Science, the human and primate atlas grant project also includes sub-projects led by researchers from 17 other institutions in the U.S., Europe and Japan.
“We are aiming to create something transformative for the field that can only be done collaboratively, by bringing in an all-star cast of experts from a variety of disciplines,” Lein said. “This is critical work: We need to understand the human brain better if we hope to treat diseases of the brain, and specifically we need a better understanding of brain function and structure. The cell atlases we’re building with the support of the BRAIN Initiative promise to lead to a more rapid understanding of the basis of many brain diseases.”

Four additional BICAN grants to the Allen Institute will support coordination and administration of the network; construction of a web-based knowledge platform using BICAN findings to accelerate scientific discovery; mapping the developing mouse brain; and tying mouse brain cells and function. Like all Allen Institute resources, the findings, techniques and data from these projects will be made publicly available.
“We’re honored to be recognized through these grants and are thrilled to partner with leading researchers around the world to build the first comprehensive map of the human brain,” said Rui Costa, D.V.M., Ph.D., President and CEO of the Allen Institute. “This atlas will open new doors in neuroscience and, ultimately, in medicine as well.”
From mice to us
These projects build off earlier NIH BRAIN Initiative-funded projects at the Allen Institute and elsewhere to map the cells of the entire mouse brain and parts of the human brain using the full suite of genes switched on in individual cells, a technique known as single-cell transcriptomics. Understanding the cell types in commonly studied mammals like mice is key to improving translational research to ultimately benefit people suffering from brain diseases and disorders, said Zeng, who is also co-leading a BICAN project headquartered at Harvard University to map cell types throughout the development of the mouse brain.
“We need to better understand the similarities and differences between the human brain and those of other primates and rodents, which often serve as the subjects for research,” Zeng said. “Comparing cell types is one of the most robust and accurate ways of comparing the brains of different species, and we are doing this at a refined level and at the most comprehensive scale we can achieve. Better relating animal studies and animal models with the structure and function of the human brain itself — that’s one of the major goals we want to achieve.”
Together, the five awards total over $173 million in funding to support these projects, including portions of the projects carried out at collaborating institutions, over the next five years.
“With the announcement of the BICAN awards, we are making an exciting transition in the overall BRAIN Initiative cell census program, which began in 2014,” said Dr. John Ngai, Director of the NIH BRAIN Initiative. “These awards will enable researchers to explore the multifaceted characteristics of the more than 200 billion neurons and non-neuronal cells in the human brain at unprecedented detail and scale — a feat in advanced technologies and cross-team research collaboration that will reveal new paradigms for understanding how pathological changes in particular groups of brain cells could cause neurologic and neuropsychiatric disorders.”
We have hundreds of different kinds of brain cells — why?
Two of the newly funded BICAN projects aim to address a key unanswered question in neuroscience: What do our different kinds of brain cells actually do?
One of these projects, led by Anton Arkhipov, Ph.D., Associate Investigator in the Allen Institute’s MindScope Program, and Marina Garrett, Ph.D., Assistant Investigator in the MindScope Program, aims to marry neuron function with cell type in the regions of the mouse brain that process what the animal sees. The scientists will develop new methods that use a high-resolution microscope to visualize specific kinds of cells in the mouse brain; these experiments will follow experiments capturing those same neurons’ activity as animals perceive and react to their visual environments.
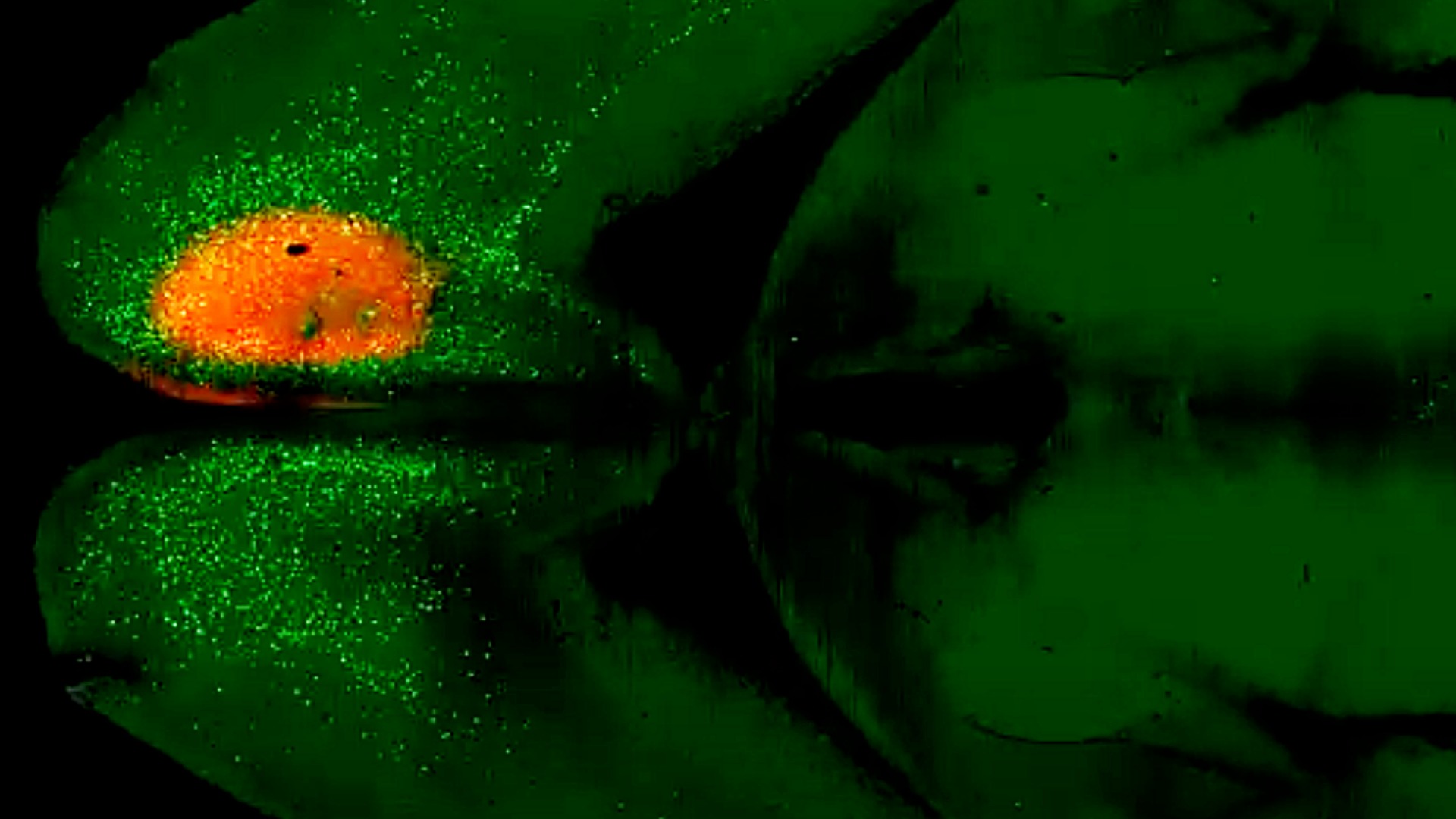
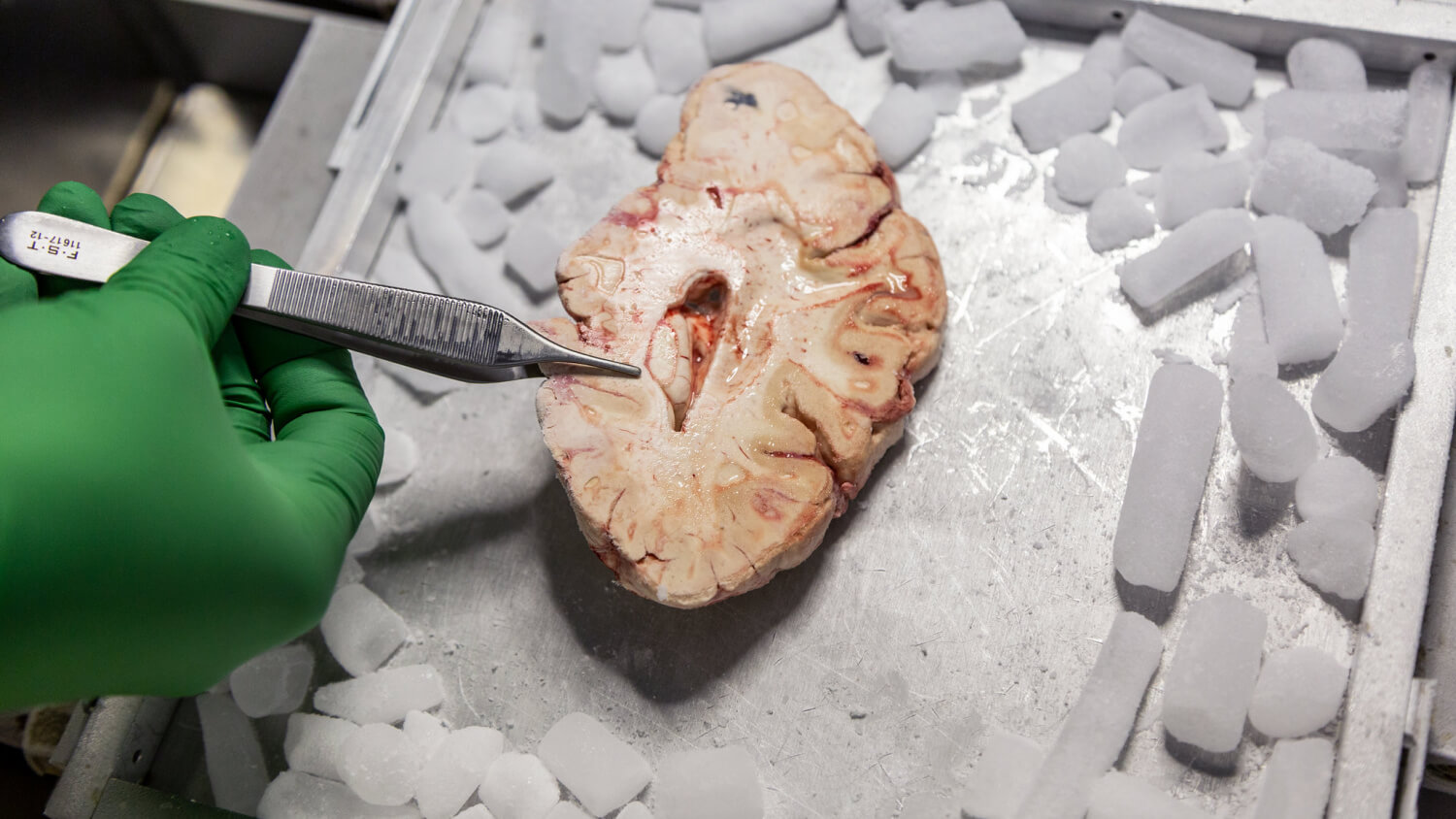
The primate brain atlas project will also marry cellular maps with brain function, using a technique known as functional MRI that captures blood flow in the brain: Active regions of the brain are correlated with higher levels of blood flow. The researchers will then correlate active regions of the brain with specific cell types using an emerging method known as spatial transcriptomics, which labels cell types in their original location in the brain. The human atlas researchers will also expand experiments long underway at the Allen Institute that study the details of living human neurons using brain tissue donated by Seattle-area brain surgery patients.
Two other awards to the Allen Institute will support coordination and knowledge sharing for the BICAN community. One team, led by Shoaib Mufti, Senior Director of Data and Technology at the Allen Institute for Brain Science, Michael Hawrylycz, Ph.D., Investigator, and Lydia Ng, Ph.D., Investigator, will develop a public, web-based knowledge platform to bring together and disseminate all the data and findings generated through BICAN work. The other group, led by Hawrylycz and Carol Thompson, Ph.D., Associate Director of Data Management, will build an engagement, coordination and outreach center for all BICAN-funded research.
“We’re thrilled to be part of this extraordinary endeavor of the BRAIN Initiative, which is building on the success we and others have had in defining brain cell types,” Zeng said. “This grant package demonstrates the leading role the Allen Institute is playing in the field — from data generation to computational analysis, as well as organization of the data, knowledge, and the entire community. We look forward to working together with our colleagues to make ground-breaking discoveries about the human brain.”
Other investigators on the BICAN grants described above include:
Trygve Bakken, Jennie Close, Song-Lin Ding, Rebecca Hodge, Tim Jarsky, Brian Kalmbach, Michael Kunst, Brian Lee, Brian Long, Jeremy Miller, Kim Smith, Staci Sorensen, Susan Sunkin, Bosiljka Tasic, Jonathan Ting, Jack Waters, and Zizhen Yao of the Allen Institute; Tim Brown, Kimiko Domoto-Reilly, Tom Grabowski, Dirk Keene, and Christine MacDonald of UW Medicine; Bruce Fischl and Eugenio Iglesias of Massachusetts General Hospital; Guoping Feng, Satrajit Ghosh, and Nancy Kanwisher of Massachusetts Institute of Technology; Winrich Freiwald of Rockefeller University; Matt Glasser and David Van Essen of Washington University School of Medicine in St. Louis; Henry Kennedy of INSERM and the Université Claude-Bernard Lyon; Takuya Hayashi of RIKEN; Fenna Krienen of Princeton University; Noah Snyder-Mackler of Arizona State University; Michael Platt of the University of Pennsylvania; Sten Linnarsson and Rickard Sandberg of Karolinska Institutet; Xiaolong Jiang and Andreas Tolias of Baylor College of Medicine; Philipp Berens of the University of Tubingen; Gabor Tamas of the University of Szeged; Christiaan de Kock and Huib Mansvelder of Vrije Universiteit Amsterdam; Boudewijn Lelieveldt of Leiden University Medical Center; David Osumi-Sutherland of EMBL-EBI; Maryann Martone and Anita Bandrowski of the University of California, San Diego; Jerry Chen of Boston University; Paola Arlotta of Harvard University; Tomasz Nowakowski of the University of California, San Francisco; and Sami Farhi of the Broad Institute of MIT and Harvard.
Research reported in this publication is supported by the National Institutes of Health BRAIN Initiative under Award Numbers UM1Mh430981, U01Mh430907, U01Mh430962, U24Mh430918, and U24Mh430919. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Citations
about the allen institute
The Allen Institute is an independent, 501(c)(3) nonprofit research organization founded by philanthropist and visionary, the late Paul G. Allen. The Allen Institute is dedicated to answering some of the biggest questions in bioscience and accelerating research worldwide. The Institute is a recognized leader in large-scale research with a commitment to an open science model. For more information, visit alleninstitute.org.