in this article
Nine years. A technique involving eight tiny electrodes. More than 1,700 connections between neurons assessed and characterized.
A team of neuroscientists at the Allen Institute has completed an incredibly ambitious undertaking to catalog the chemical connections, or synapses, between thousands of neurons in mouse and human brains.
The result: a much more complete picture of the diversity of mammalian synapses, the routes by which information is passed from neuron to neuron in the brain. The team published a study describing their results in the journal Science today and released their synapse data publicly online.
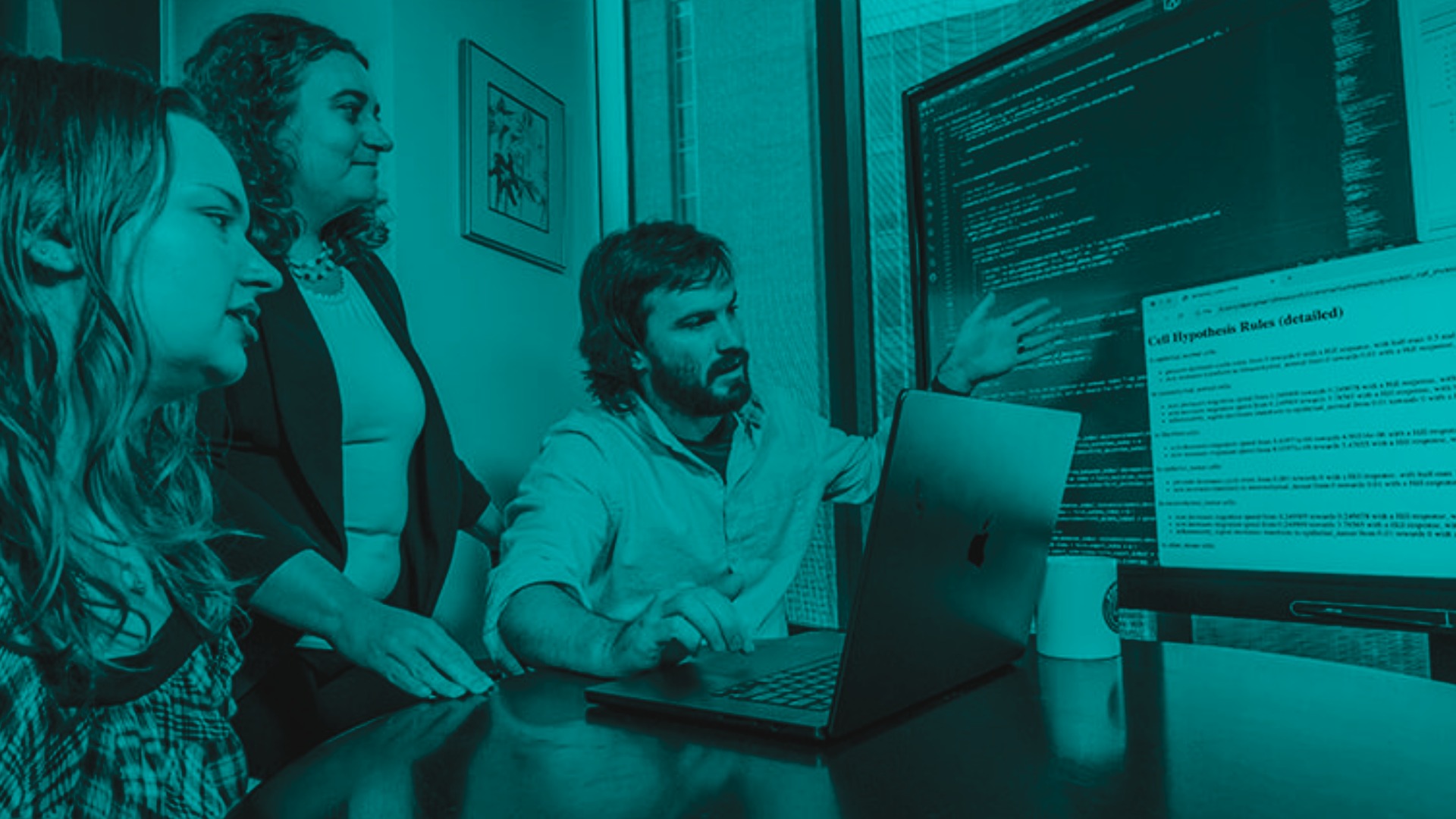
“At the Allen Institute, we spend a lot of time thinking about all the different types of cells in the brain. This project looked at those different cell types and asked which ones are connected to each other and what those connections look like,” said Luke Campagnola, Ph.D., a scientist at the Allen Institute for Brain Science, a division of the Allen Institute, who led the study together with scientist Stephanie Seeman, Ph.D., and Tim Jarsky, Ph.D., Associate Director of Electrophysiology. “It’s a huge cataloging project with the idea that the more we know about the structure of the brain, the more likely we are to be able to understand it in the future.”
Eavesdropping on neurons’ chatter
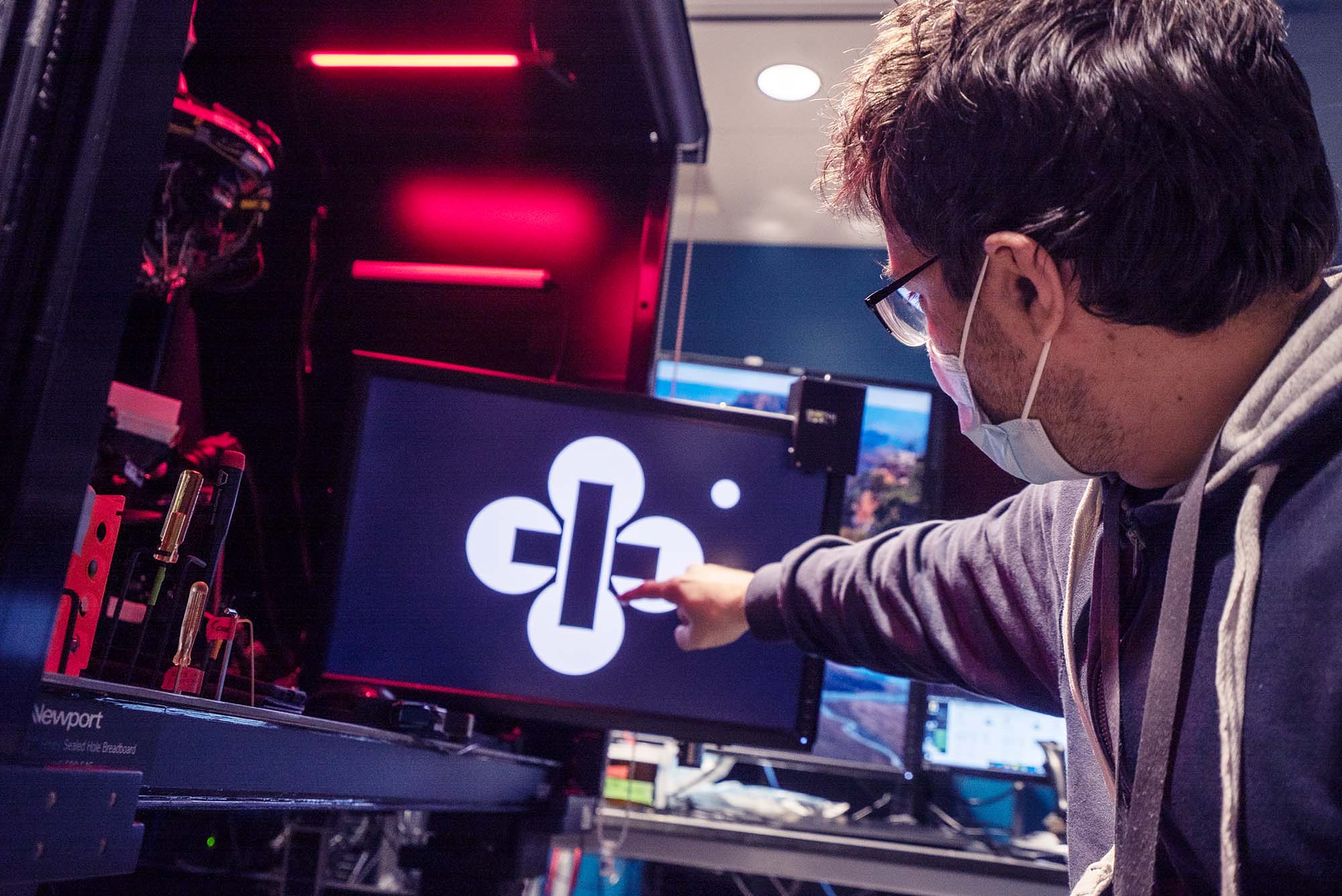
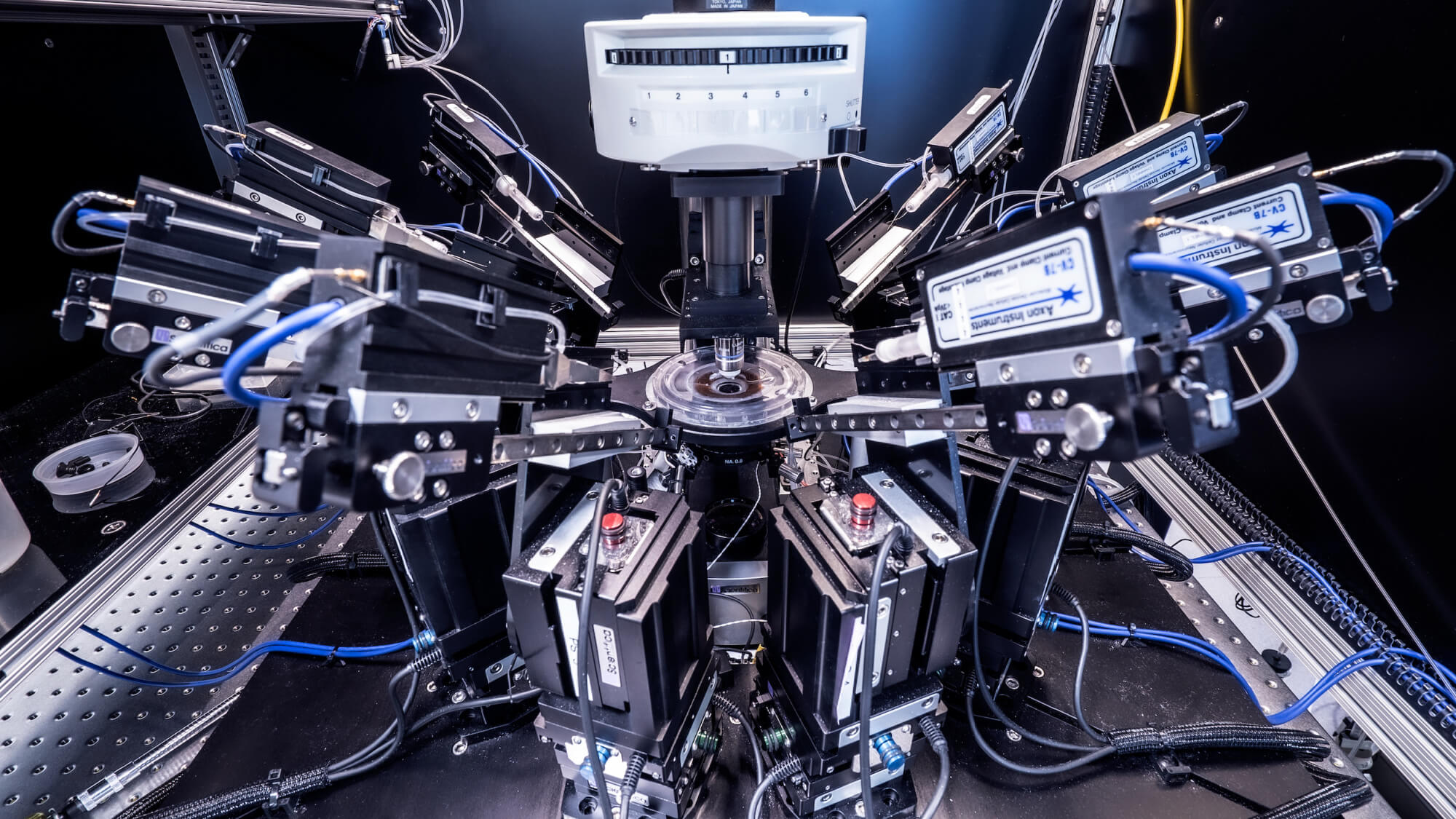
While there are many techniques that can reveal the pathways information travels in the brain, there are fewer methods to study connections between cells in detail. The method the Allen Institute team used is known as “multipatch,” and it involves piercing eight neighboring neurons with tiny electrodes to eavesdrop on their electrical signals, using a specially built microscope rigged up with eight robotic arms to hold the electrodes.
You can think of the pathway of a thought or any other information in the brain as a complicated — and really fast — relay race. Electrical signals travel the length of a single neuron, from its round central body down a long, specialized tendril called an axon, until that axon reaches a connection point with another neuron. The electrical signal triggers a chemical signal, where molecules known as neurotransmitters flow from one cell to the next and serve as the signal hand-off, beginning a new electrical current in the next cell.
Synapses, the points where the two neurons meet and where the chemical exchange happens, aren’t one-size-fits-all. Like the cells they connect, synapses have natural variability, and neuroscientists want to understand the differences between different types of synapses as well as which cells are likely to talk to each other. The multipatch method can get at both questions, a few cells at a time. To analyze the details of more than 1,700 different synapses, the scientists performed experiments on nine different classes of neurons, assessing nearly 24,000 potential connections. These data were collected in under three years, but the project had its inception at the Allen Institute in 2013 — it took several years for the scientists to troubleshoot and ramp up to large-scale data collection.
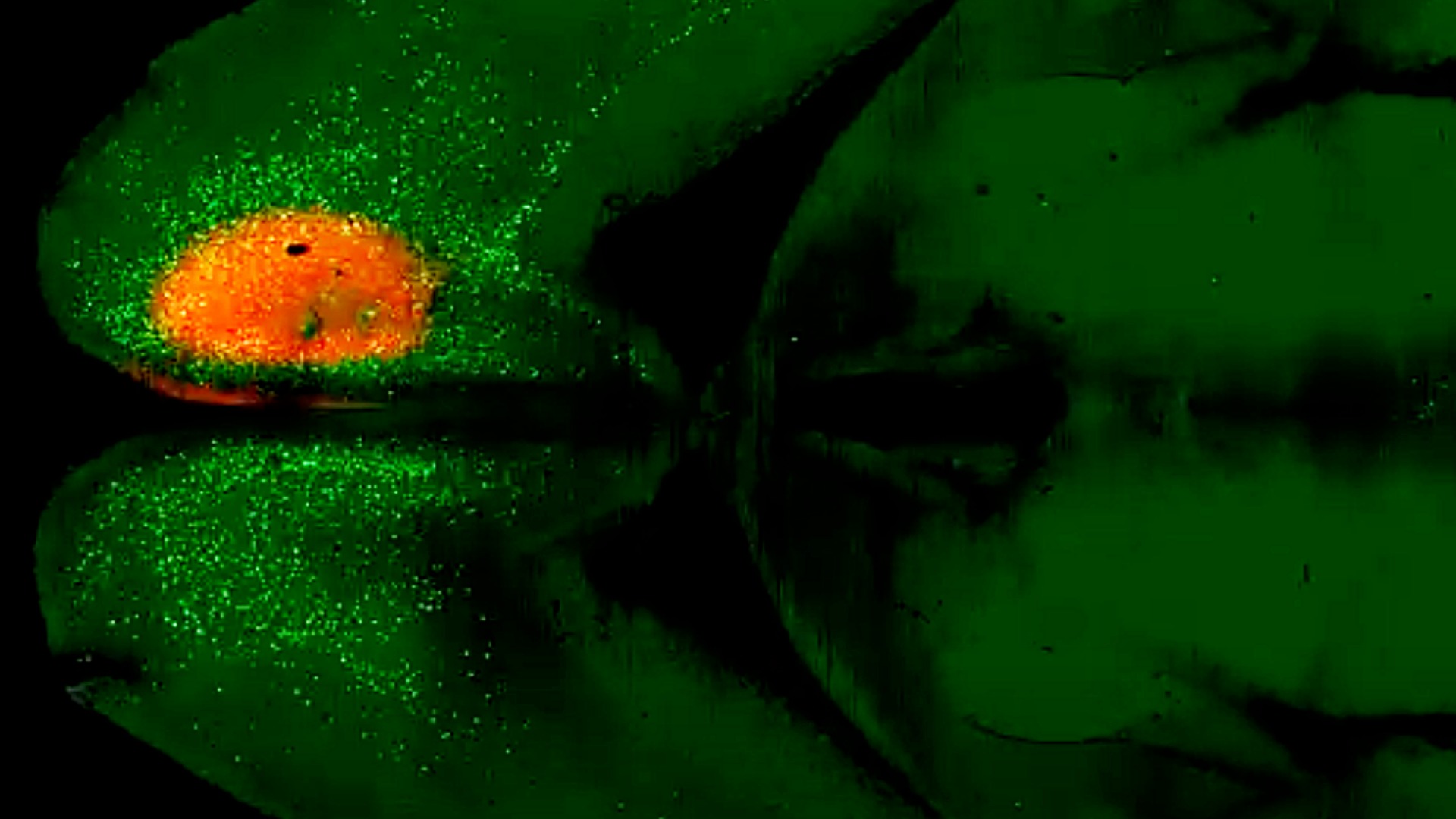
Most of these experiments involved visual neurons in the region of the mouse cortex, the outer layer of the brain, that processes visual information. About 200 synapses in the dataset come from human neurons, capturing the electrical signals in still-living pieces of human brain tissue donated by patients undergoing brain surgeries in Seattle-area hospitals. In these surgeries, a small piece of healthy tissue may be removed to reach the diseased part of the brain; with the patient’s consent, this healthy tissue is kept alive and transported to the Allen Institute where scientists can glean important information from living human brain cells.
Studying tiny voltage changes
To produce such a large and standardized dataset, the scientists automated as much of the multipatch experiments as they could. Still, the method is tricky and sometimes tedious. Experimenters need to identify the neurons of interest and find eight healthy looking cells in close proximity, and then guide the robotic arms to those cells, finally piercing the delicate edge of each cell manually. Once all eight cells are “patched,” the experimenter jolts one neuron with a tiny current and records any signals that make it to the other seven, and then onto the next neuron until all eight have been coaxed to send signals. It’s a long process — a highly trained researcher can run maybe three successful multipatch experiments a day.
“It’s a roller coaster, doing these experiments,” Seeman said. “Some days, things are just clicking along and you get great data, and then you’ll have days of turmoil where it seems like nothing you do is working.
The electrical response of the connected cells, their voltage change, gives the scientists important clues about the nature of the connections between the cells, even without looking directly at those synapses. But the electrical changes can be difficult to parse, Jarsky said.
“The scale of these voltage changes is typically sub-millivolt, so less than one-thousandth of a volt,” he said. “These are very small signals, and that’s one of the reasons this approach is so hard and complicated.”
The scientists gather features of synapses that include their strength, their speed, and their variability or plasticity (how much the same synapses change their signaling over the course of an experiment), as well as whether two given cells have a synapse together in the first place. Not all neighboring cells are connected, but cells that are close are more likely to connect than those that are far away, even though some neurons do connect across very long distances in the brain.
While scientists could sift through the data for insights on particular neurons of interest, the researchers think their open-access data will be most useful to inform better computational models of how neurons connect into circuits that allow our brains to do what they do. Sophisticated and realistic models of the brain could incorporate details not only about the diversity of different neurons, but the diversity of synapses as well.
“Modeling is incredibly important, because the brain is too complex a system to use your intuition to figure out how cells should be behaving,” Jarsky said. “Even with the subset of kinds of neurons in this dataset, we have so many possible interactions with many different types of signaling. You really need a model to understand how the circuit could behave when you put all those elements together.”
Research described in this article was partially supported by awards from the National Institutes of Health, including award R01DA036909 from the National Institute on Drug Abuse and awards U19MH114830, U01MH114812 and RF1MH121274 from the National Institute of Mental Health. The content is solely the responsibility of the authors and does not necessarily represent the official views of NIH and its subsidiary institutes.
Citations
about the allen institute
The Allen Institute is an independent, 501(c)(3) nonprofit research organization founded by philanthropist and visionary, the late Paul G. Allen. The Allen Institute is dedicated to answering some of the biggest questions in bioscience and accelerating research worldwide. The Institute is a recognized leader in large-scale research with a commitment to an open science model. For more information, visit alleninstitute.org.