in this article
authors
The power of prediction is everywhere in our lives: we try to forecast the weather, swings in the stock market, technological advancements, and much more. That’s also true in science.
In biology, it’s hard enough to understand how cells interact with each other. It’s even more complex to predict how these cells will change over time as they do. New technology can measure genes and proteins in individual cells, but these methods provide just a snapshot, leaving unanswered questions like how cells eventually become cancerous or immune-resistant. Meanwhile, mathematical and computer models can simulate how cells evolve, but they’re hard to build and require advanced programming skills. However, a new study combining biological research and computational modeling could democratize this field, helping scientists forecast how cells change, with ramifications for cancer research, immunology, and other areas.
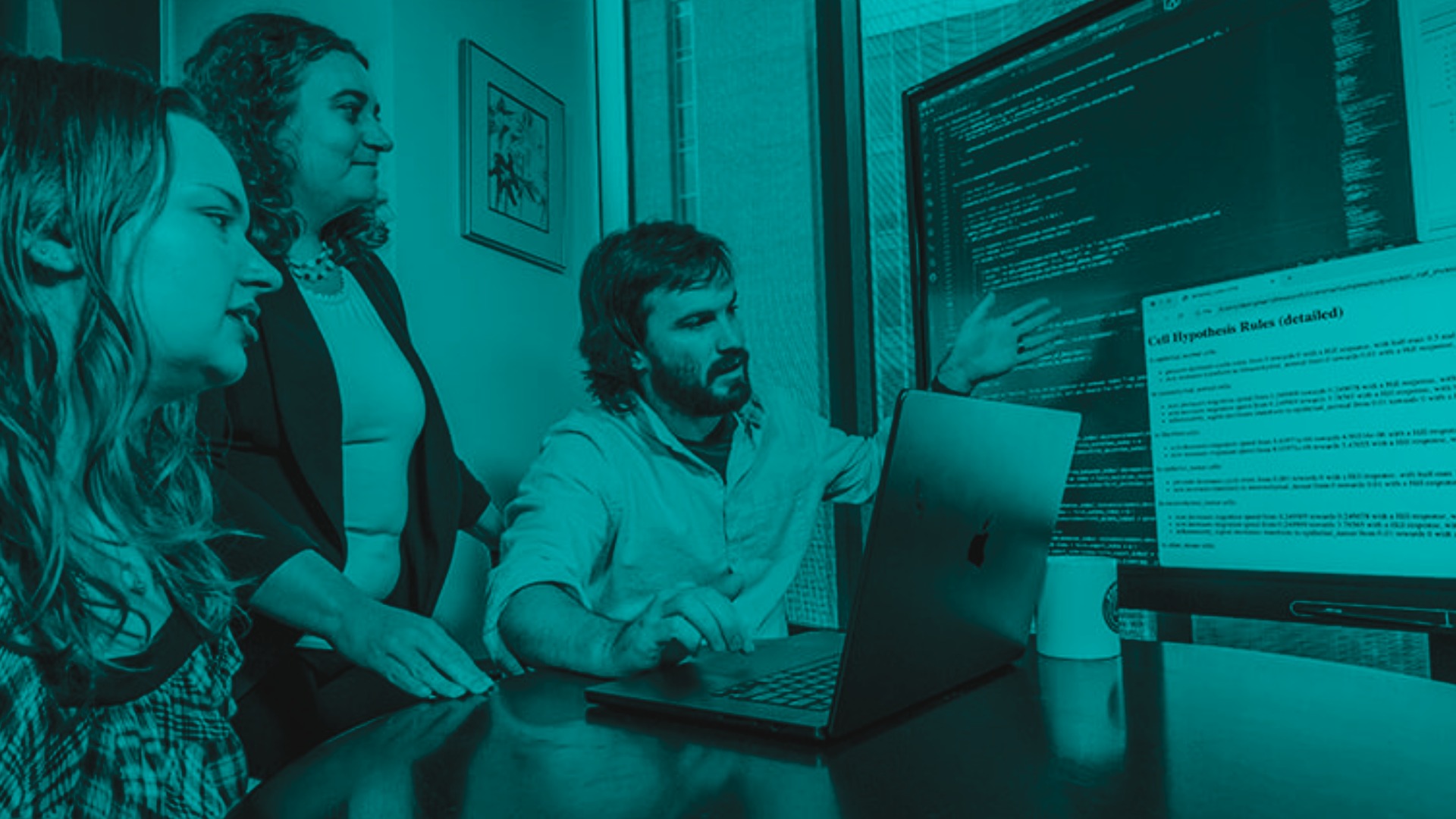
A team of scientists from the University of Maryland and Johns Hopkins University developed an understandable modeling language that allows researchers to describe, in words and sentences, how cells behave and respond to their environment. This cell behavior grammar enables them to plug intuitive statements into the program, such as “oxygen decreases cell death”, and run virtual experiments on tissues such as tumors.

We can take these snapshots and put them into this virtual cell laboratory and basically hit ‘play’. Then we watch the movie unfold and make predictions on the hypothesis based off what we see.
- Genevieve Stein-O’Brien, Ph.D., assistant professor of Neuroscience and Neurology at Johns Hopkins School of Medicine
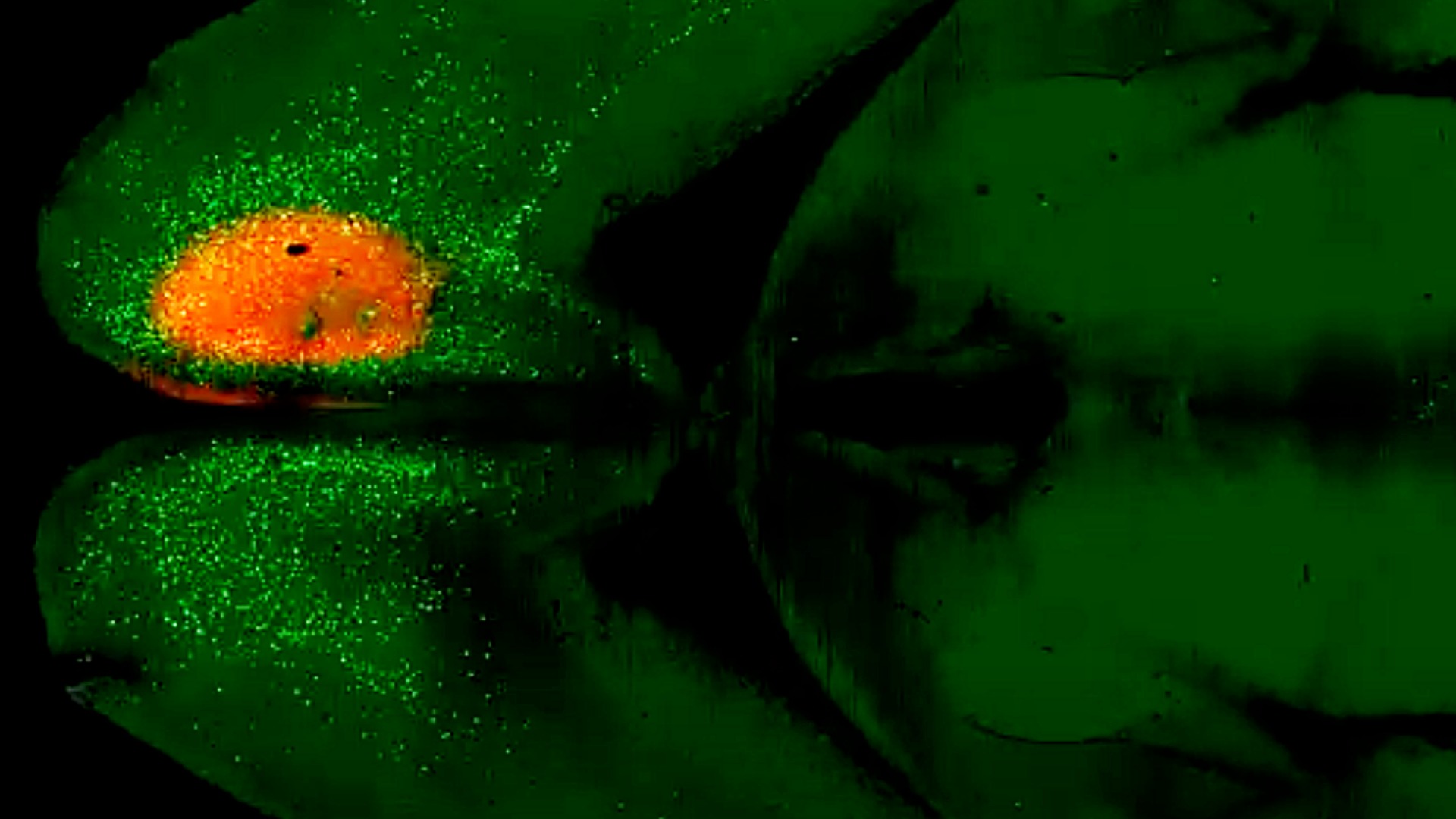
One important finding involved simulations of pancreatic cancer using genomic data from human patients. The models predicted that signals from immune cells promote invasion primarily by increasing cancer cell movement—known as motility—rather than cell division, which they later confirmed with further experiments. In the future, they hope to study other cancers.
“We’re trying now to extend this modeling framework to see if we can predict the development of pancreatic cancer and learn which lesions will grow and which ones won’t,” said Elana Fertig, Ph.D, Director of the Institute for Genome Sciences and Dean Albert E Reece Endowed Professor of Medicine and Epidemiology at the University of Maryland School of Medicine.
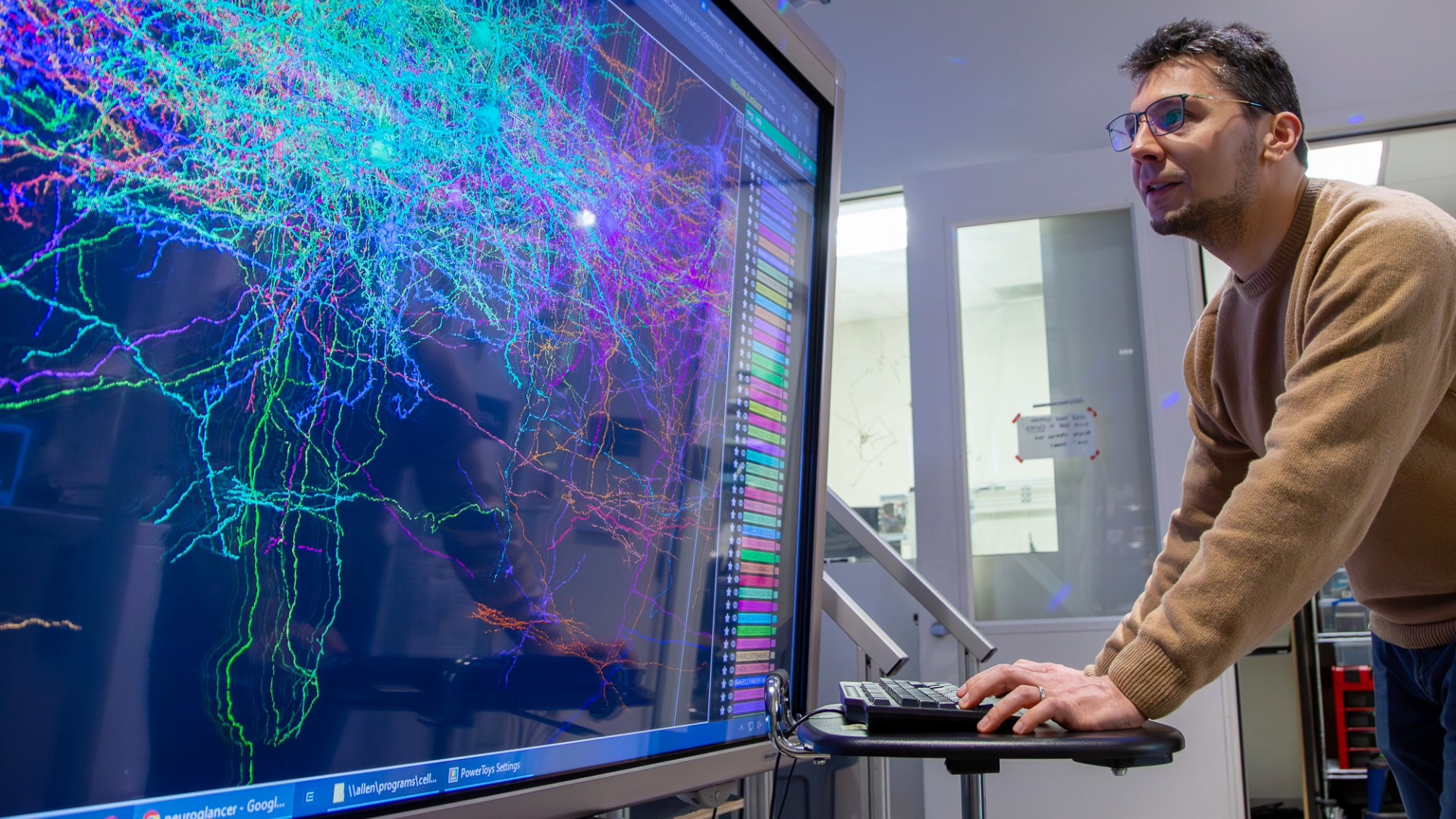
To test the model and show how broadly it could be applied, researchers used the Allen Institute’s Mouse Brain Atlas to simulate cell layer formation during brain development particularly in the cortex.
“The ability to load a whole brain of agents that are potentially connected and interacting with each other is extremely powerful,” said Stein-O’Brien.
“Agent-based modeling is a type of mathematical modeling that needs a lot of data support because we want to represent individual cells as discrete software agents that interact with one another and with their environment,” said Daniel Bergman, Ph.D., assistant professor at the Institute for Genome Sciences at University of Maryland School of Medicine and assistant professor of Pharmacology and Physiology. “So I like to think of it as the truest digitalization of how we can see cells in biology.”
The software is open source, so other scientists can access it for their research without requiring a deep understanding of computer programming. This work could eventually lead to computer programs that help treat cancer patients by creating a “digital twin” of the patient, which could help predict the best course of treatment. Scientists hope this technology will provide a bridge to a predictive future that largely hasn’t existed before in such an accessible way.
Citations
about the allen institute
Allen Institute is a 501(c)(3) nonprofit medical research organization dedicated to accelerating science for a healthier world. Through large-scale, multidisciplinary research initiatives, the Institute generates foundational knowledge, data, tools, and models that are shared openly with the world to advance our understanding of life and health. Founded by Jody Allen and the late Paul G. Allen, Allen Institute is supported primarily by the Fund for Science and Technology.