in this article

By Rachel Tompa, Ph.D. / Allen Institute
Scientists have built a new space for collaboration — at the microscopic level.
The Allen Institute for Cell Science’s Simularium Viewer, described today in a publication in the journal Nature Methods, aims to bring together disparate views of cells and molecules and to enable visualization of computational renderings of living systems.
The viewer is a web-based platform where users can share their own data or computational models to generate visualizations, which can help scientists better interpret their data and more easily share their findings with others.
“The goal is to make it easier for experimental biologists to collaborate with computational biologists who are working on biological simulations,” said Blair Lyons, Simulation Software Design Engineer at the Allen Institute for Cell Science who led the development of Simularium along with Graham Johnson, Ph.D., Senior Director of the Animated Cell team, and Eric Isaac, a former software engineer at the Allen Institute for Cell Science.
Simularium aims to address a number of problems in computational cell biology by allowing users to see the outputs of their computational models, by integrating different models of the same system (for example, to visualize the same cellular structure at different levels of magnification), and by allowing non-experts to use and understand the outputs of computational models.
Computational models — virtual simulations of some real-life process, usually simplified — can be hugely helpful in predicting and better understanding cell biology. But most models are difficult for non-experts to interact with, let alone understand, and because models are usually made for one specific purpose, their output isn’t always comparable to other models.
“A lot of computational biology approaches — just like in any other scientific specialty — can be esoteric, or at least require special skills to use,” Johnson said. “The types of models that Simularium supports can be visualized just a mouse-click away.”
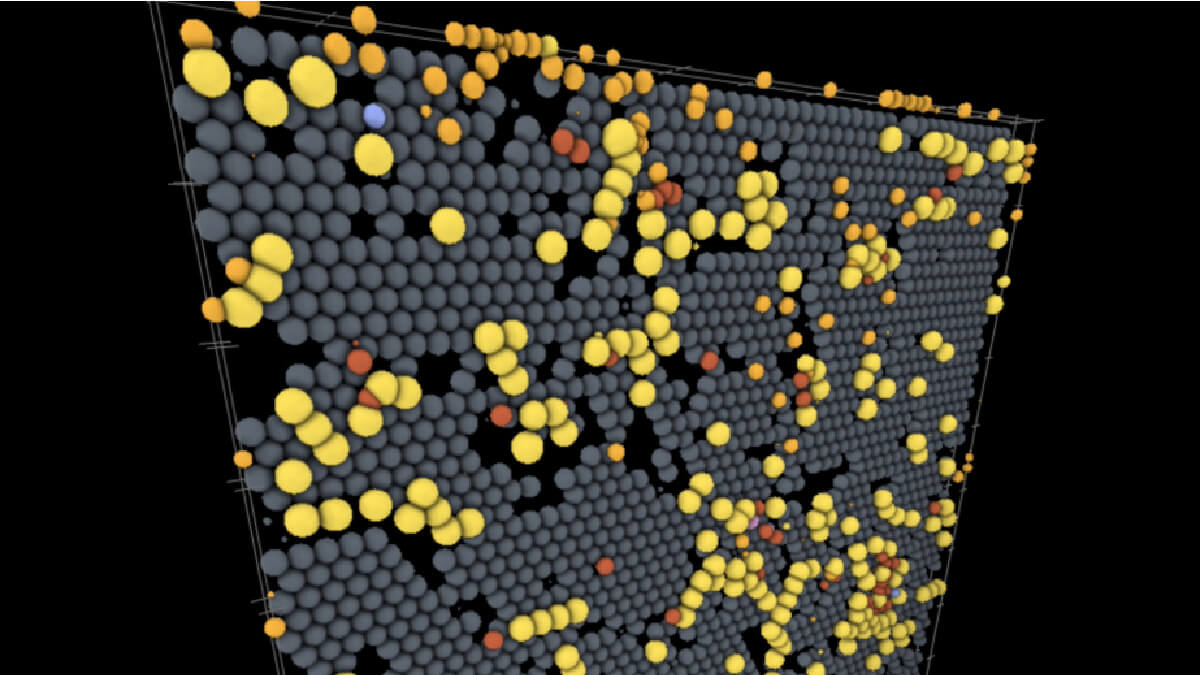
Some computational models might give results in a string of numbers, or lines of code. Simularium translates models of spatial objects into an onscreen 3D visualization and related 2D graphs that aid interpretation. The viewer also allows users to zoom in and out, show or hide parts of the model, and rotate views.
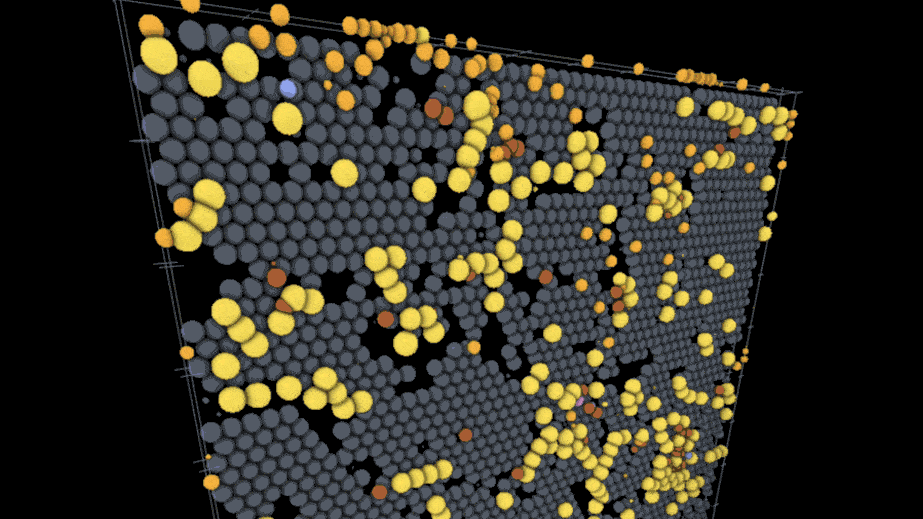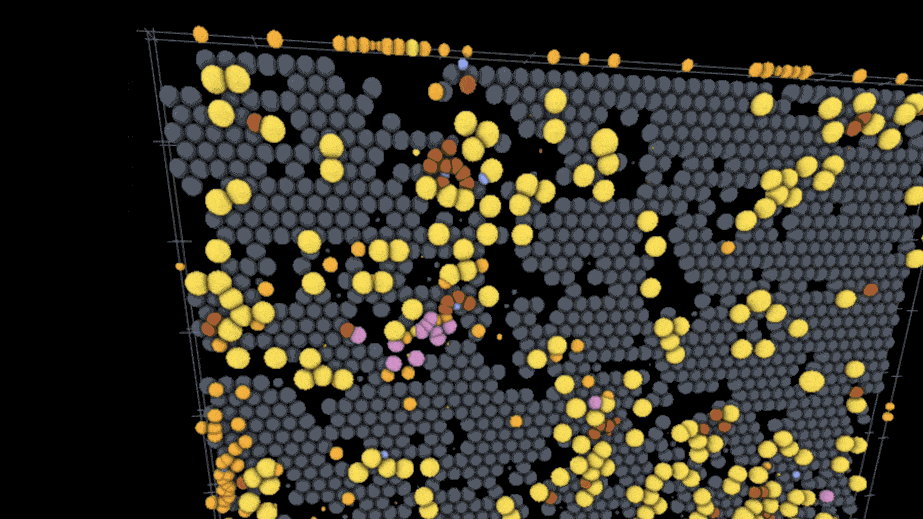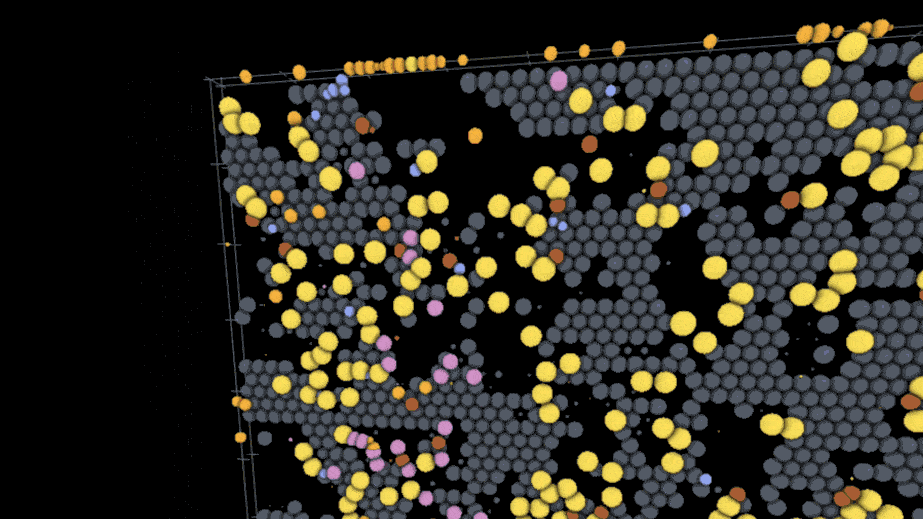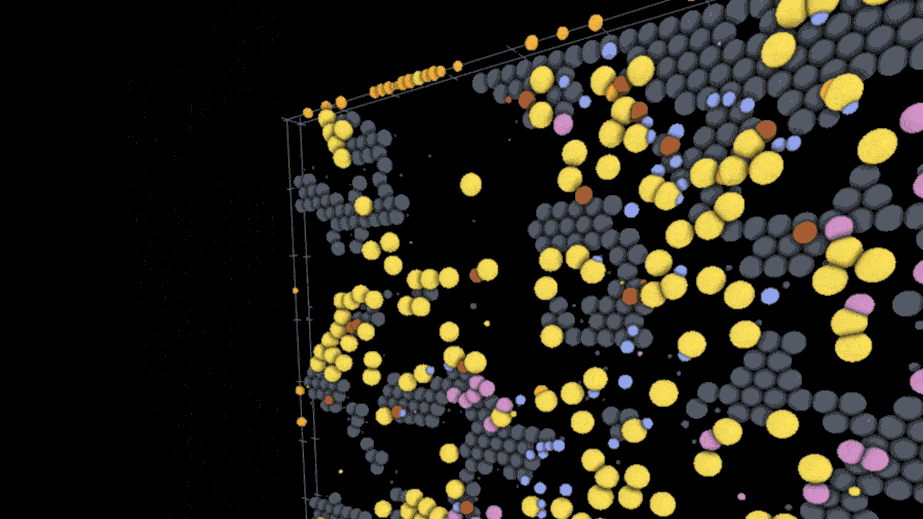
One example visualization on the site models the interactions of SARS-CoV-2 virus particles with different kinds of immune cells on the lining of the lung using models and data from an external team of researchers; users can play through the time course of the model to watch the infection spread while rotating the surface of the tissue and hiding or showing the different cell types involved. Another visualization created by a different lab deals with action at a much smaller scale — a lipid membrane wrapping a nanoparticle.
In addition to being a platform for exploration and collaboration, Simularium’s developers foresee it as complementing how scientific findings are shared with the community.
“Simularium could expand what is available for scientific publishing,” Isaac said. “We hope it will become a way to make interactivity and web-native sharing extremely easy to integrate alongside scientific findings with direct access to the scientific data.”
Citations
about the allen institute
The Allen Institute is an independent, 501(c)(3) nonprofit research organization founded by philanthropist and visionary, the late Paul G. Allen. The Allen Institute is dedicated to answering some of the biggest questions in bioscience and accelerating research worldwide. The Institute is a recognized leader in large-scale research with a commitment to an open science model. For more information, visit alleninstitute.org.