events
enjoying respectful, inclusive events
Thank you for participating in an Allen Institute event.
Events are central to our mission of scientific discovery and our commitment to big, team, and open science. We are dedicated to creating environments—both in person and virtual—where all participants feel safe, respected, and welcomed.
All attendees, speakers, organizers, and staff are expected to uphold this Code of Conduct and contribute to a positive, inclusive experience for everyone.
Events are central to our mission of scientific discovery and our commitment to big, team, and open science. We are dedicated to creating environments—both in person and virtual—where all participants feel safe, respected, and welcomed.
All attendees, speakers, organizers, and staff are expected to uphold this Code of Conduct and contribute to a positive, inclusive experience for everyone.

Summer Workshop on the Dynamic Brain 2026
The 2026 Summer Workshop on the Dynamic Brain is an intensive, project-based residential course with a focus on the neurobiology of sensory processing, coding, and neural population dynamics.
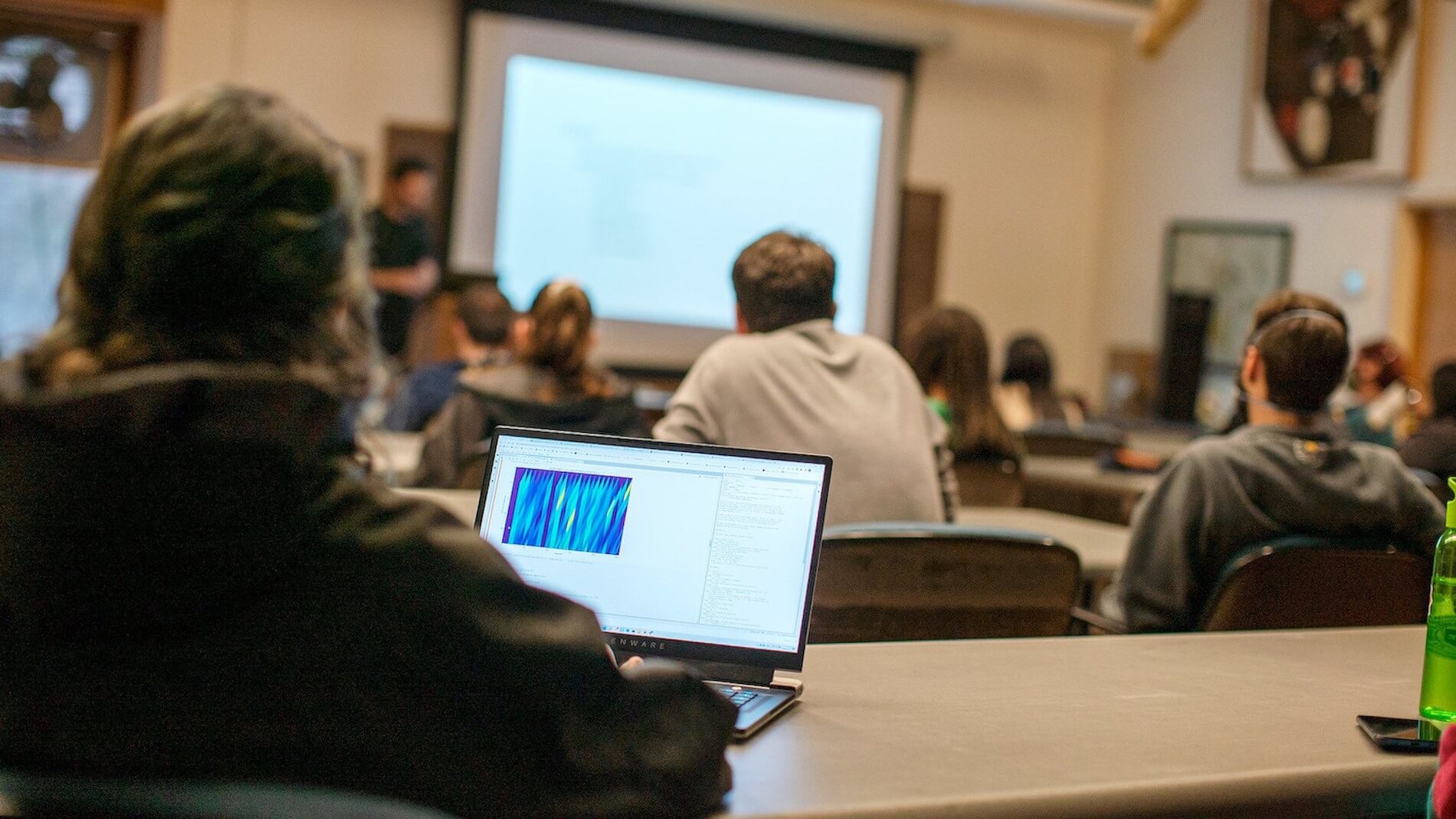

explore all events
Thank you! Your submission has been received!
Oops! Something went wrong while submitting the form.

2026 Computational Cell Types Workshop
In this hackathon-style workshop, attendees will be introduced to single cell- and spatial transcriptomic datasets from the Allen Institute and their collaborators, while working in teams to use these datasets for data analysis, tool creation, model development, and more.
Brain Science
Graduate
Postdocs
Scientists

2026 Cascadia Advanced Genomic Technologies
The Cascadia Advanced Genomic Technologies (CAGT) Meeting is a growing annual conference that brings together Cascadia’s (Oregon, Washington, and British Columbia) leading minds to accelerate discovery at the intersection of biology and technology.
SeattleHub
Early Career Scientists
Graduate
Postdocs
Scientists

Implementing Open Neuroscience Teaching Tools: Post-FUN Pedagogy Workshop
Join the Allen Institute for the "Implementing Open Neuroscience Teaching Tools" workshop at North Central College in Naperville, IL on July 19th, 2026.
Education and Engagement
Workshop
Education
Educators

2026 Allen Institute Modeling Software Workshop
Join us at the 2026 Allen Institute Modeling Software Workshop for a hands-on opportunity to learn how to use open-source software modeling tools developed by the Allen Institute and our collaborators at the University of Illinois Urbana-Champaign (UIUC).
Brain Science
Graduate
Postdocs
Scientists
Incorporating open connectomics data into teaching neuroscience
Join a community of undergraduate neuroscience educators at the Incorporating Open Connectomics data into teaching Neuroscience workshop on July 23 – July 24, 2026, hosted by the Allen Institute.
Brain Science
Education
Education and Engagement
Educators

Coding with Allen Institute Data: A Python Workshop for Undergraduate Educators 2026
This workshop will introduce undergraduate neuroscience educators to working with Allen Institute open datasets using Python.
Brain Science
Neural Dynamics
Educators

Summer Workshop on the Dynamic Brain 2026
The 2026 Summer Workshop on the Dynamic Brain is an intensive, project-based residential course with a focus on the neurobiology of sensory processing, coding, and neural population dynamics.
Brain Science
Graduate
Scientists

Cascadia Mucosal Biology Symposium
The inaugural Cascadia Mucosal Biology Symposium hosted by the University of Washington and the Allen Institute in Seattle, Washington, September 10 - 11, 2026.
Immunology
Early Career Scientists
Postdocs
Scientists
Students
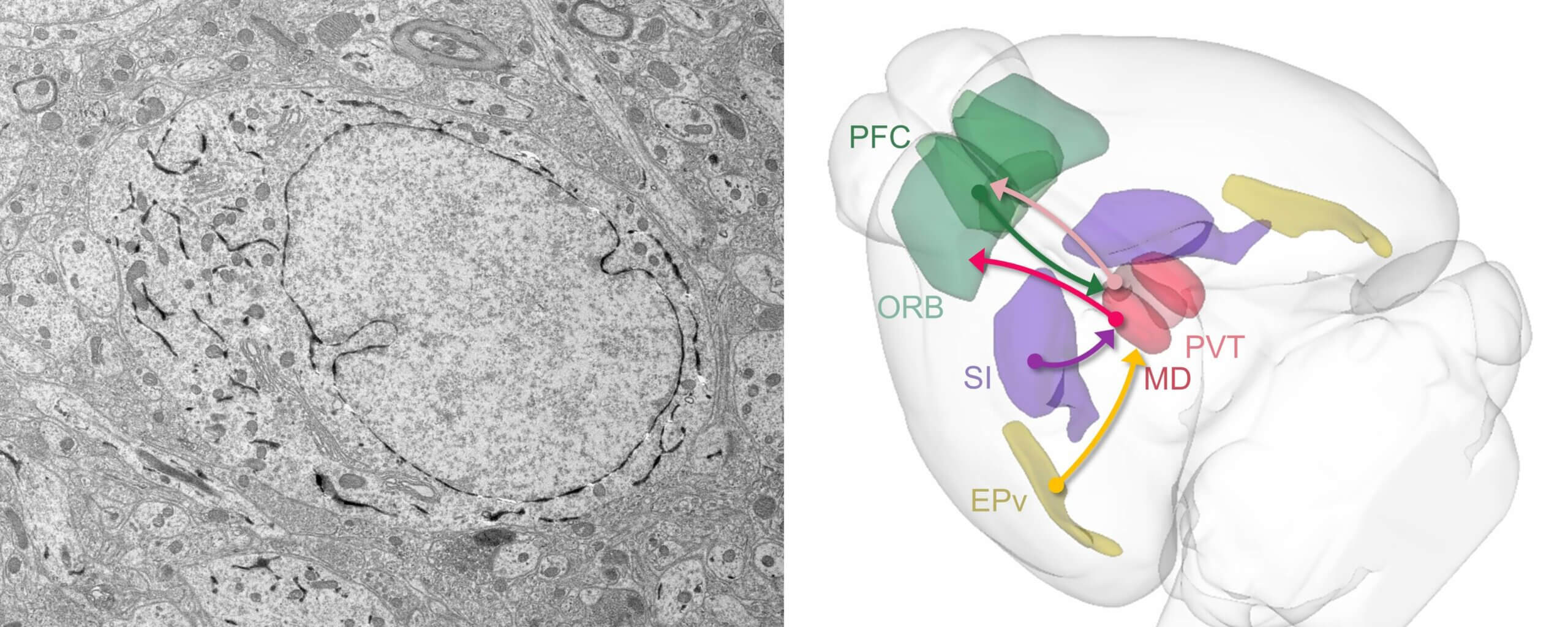
Emerging connectomics of thalamus and striatum workshop
Join us for a hands-on opportunity to learn how to use volumetric electron microscopy (EM) data and address biological questions central to the thalamus and striatum at the Emerging Connectomics of the Thalamus and Striatum Workshop, hosted at by the Allen Institute.
Neural Dynamics
Graduate
Postdocs
Scientists

Stem Cell & Developmental Biology Early Career Symposium
For the first time, the International Society for Stem Cell Research (ISSCR), the Society for Developmental Biology (SDB), and the Allen Institute for Cell Science (AICS) are collaborating to present a three-day scientific symposium led by early-career scientists.
Cell Science
Scientists
