Staci Sorensen
Associate Director, Neuroanatomy
Staci Sorensen joined the Allen Institute as a postdoc in 2008 to explore the relationship between anatomically- and genetically-defined cell types in mouse neocortex. Currently, as a Scientist II in the Mouse Cell Types group, she is investigating the range of morphologically-defined cell types in the primary visual cortex and the lateral geniculate nucleus of the thalamus. This is part of a larger effort to understand the functional components that make up the visual thalamocortical circuit. Before coming to the Allen Institute, she earned her doctoral degree in Neurobiology and Behavior working with Ed Rubel at the University of Washington. The focus of her research was to understand the role of neuronal input in the dynamic regulation of dendritic structure in the chick auditory system. She earned her B.S. in psychology at the University of Washington, and worked with Jaime Olavarria and Theresa Jones to study the effects of visual experience on the development of callosal structure in the rat visual cortex.
research focus
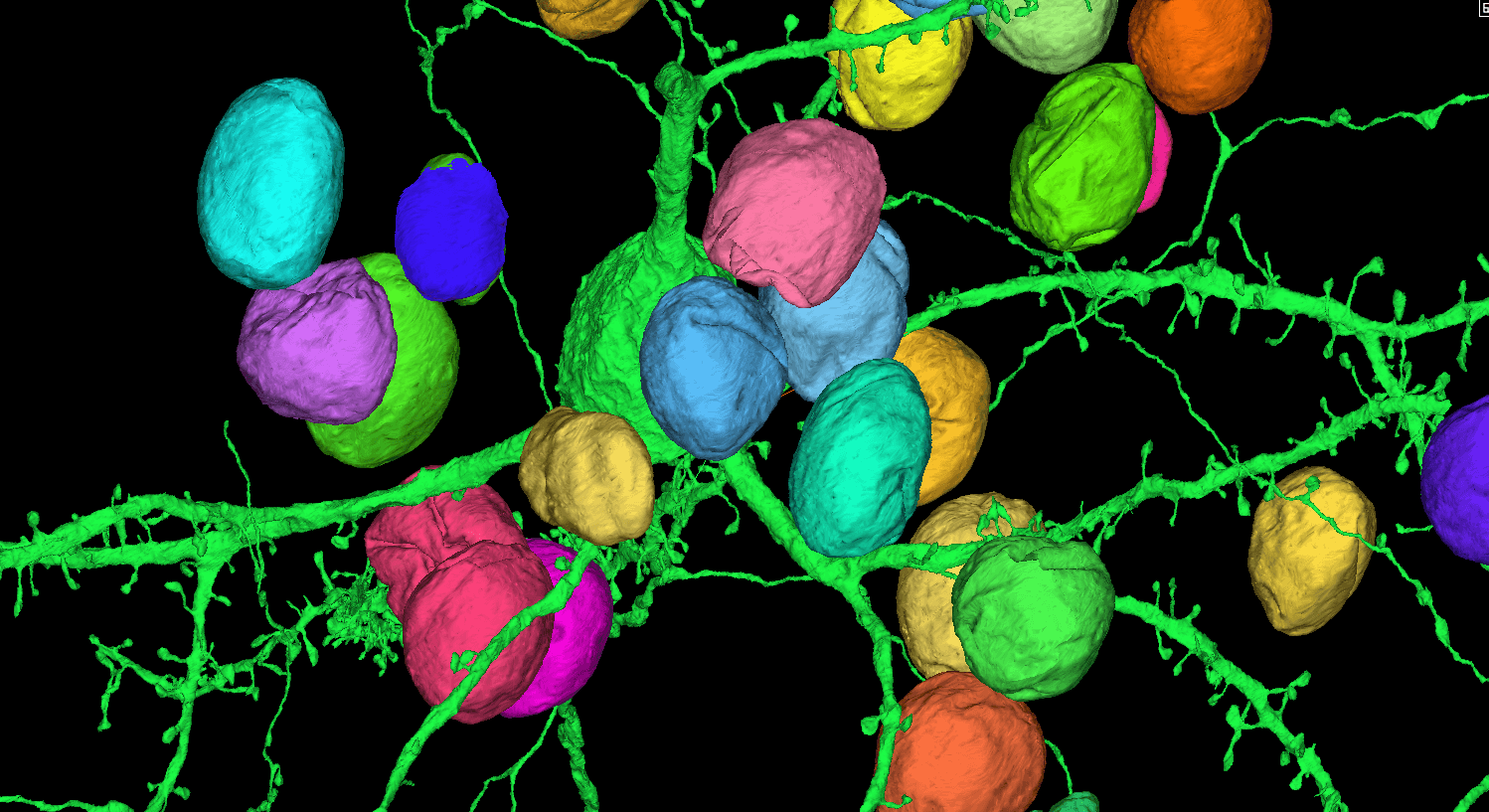
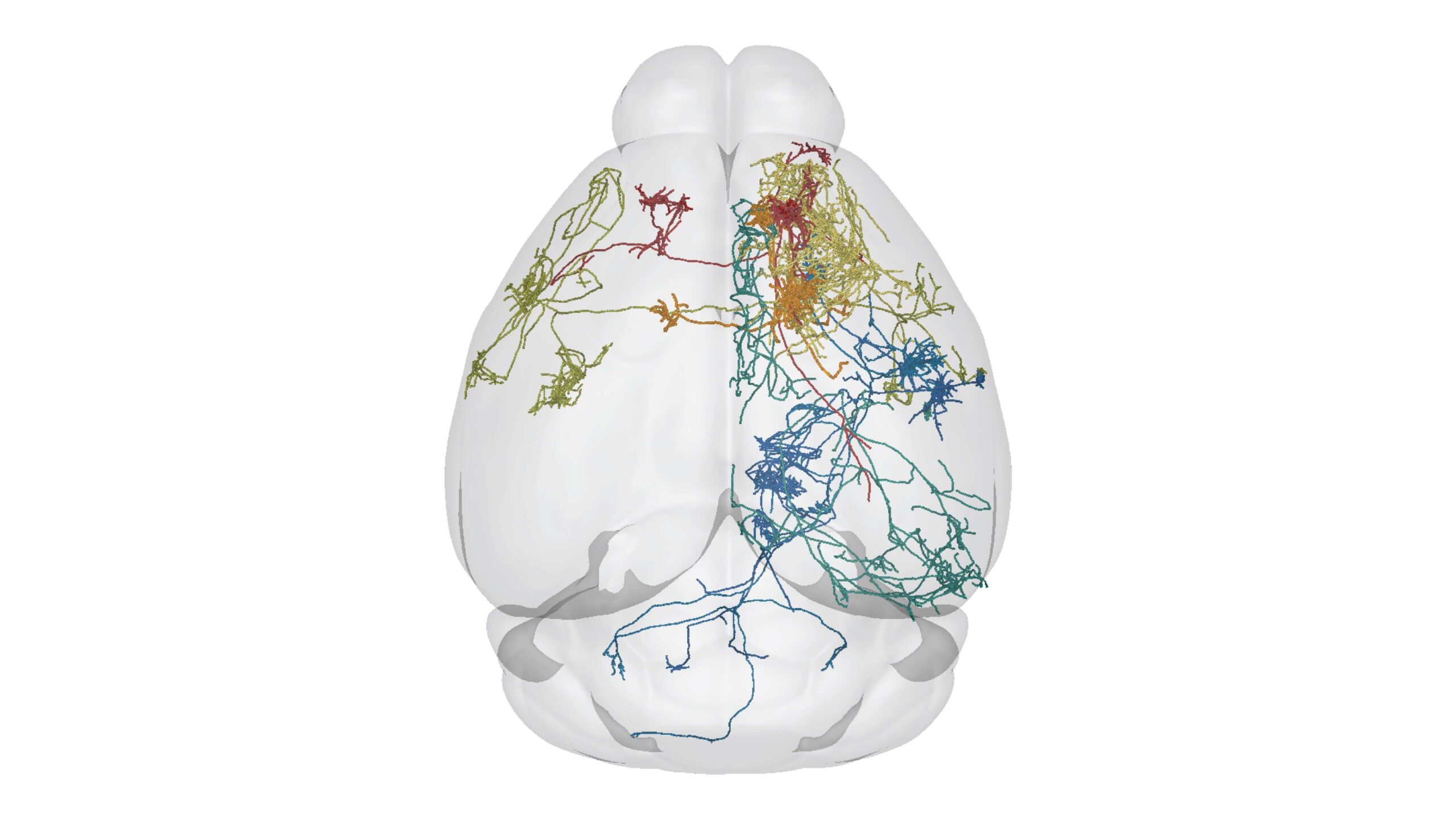
I am interested in understanding how the brain develops, how it is organized, and how its structure is regulated throughout life. Questions of particular interest to my current work relate to the organization and development of the mouse visual system and the neocortex. I am part of a large effort at the Allen Institute focused on understanding the diversity of cell types in the primary visual cortex and lateral geniculate nucleus of the thalamus. Though it is well accepted that multiple cellular properties must be used to define a cell type, my work focuses on describing cells from a morphological perspective. To do this, we use high resolution brightfield and fluorescence microscopy (multi-photon and resonant confocal) in combination with in vitro and in vivo single cell labeling methods. From montaged, multi-plane 2D images we generate a 3D structural description of individual neurons. These descriptions will be used in combination with other data to answer questions such as: How many different cell types exist in these brain regions? What are their morphologies? Across cell types, how does morphology relate to physiology, gene expression, connectivity and function? This work will help us to better understand how the mouse visual system is organized, and also to identify the features that are essential to define a cell type. I also lead a smaller project at the Institute related to characterizing cortico-cortical axon development across neonatal time points in the mouse cortex. For this work, referred to as the Developmental Connectivity Project, multiple cortical brain regions are labeled by Cre-dependent and Cre-independent adeno-associated viruses in layer-specific transgenic mouse lines. These data will give us a better understanding of how gene-defined and anatomically-defined projection neuron classes develop relative to one another within and across brain regions. It will also complement the Allen Mouse Brain Connectivity Atlas, which already contains a large number of cortical datasets from adult wildtype and transgenic mice.